A drug resistance gene mediates stringent response in bacteria
Mycobacteria represent an important group of bacteria. Mycobacterium tuberculosis, the causative agent of tuberculosis (TB), infects about a third of the world population, of which 5% to 15% develop active disease. Furthermore, a rapid emergence of multidrug resistance (MDR), extensive drug resistance (XDR), and total drug resistance (TDR) strains of M. tuberculosis make the study of stringent response in mycobacteria an important area of research. Rifampicin (Rif) is the most effective first line drug used against TB treatment. However, extended use of Rifampicin has led to high resistance in M. tuberculosis. Rif resistance arises due to mutations in its target protein, RpoB, a component of a protein complex, RNA polymerase, which transcribes genes into RNA copies. It is desirable to screen for the newer versions of rifampicin to act on the resistant strains of M. tuberculosis. However, the pathogen, M. tuberculosis must be grown under highly contained specialized facilities. On the other hand, use of model organisms can circumvent this requirement for at least the primary screening of the compounds. M. smegmatis, a soil bacterium, which can be grown without containment facilities is often used as a model organism for M. tuberculosis.

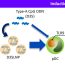
Fig. 1. Model for physiological roles of M. smegmatis Arr. A direct function of Arr is to ADP-ribosylate rifampicin (or some of its analogs, Baysarowich et al. 2008) to inactivate the antibiotic and confer rifampicin resistance. The other function of Arr is to interact with the N-terminal sequence of MsmL11 to result in production of ppGpp, the stringent response alarmone. The interaction between MsmArr and MsmL11 is species specific. To prepare the model, the structure of Arr was adapted from Baysarowich et al. 2008, and the diagrammatic sketch of ribosome highlights L11. Sketches are not drawn to scale. [Reproduced from Agrawal, P., Varada, R., Sah, S., Bhattacharyya, S. and Varshney, U. (2018) Species specific interactions of Arr with RplK mediate stringent response in bacteria. J. Bacteriol. 200, e00722-17. doi: 10.1128/JB.00722-17]
In the present work, we sought to understand the mechanism of Arr action. Reports from the other groups (Stallings et al., 2011) showed that the expression of arr is upregulated during double stranded DNA break in the genome, starvation, and genotoxic antibiotic (ciprofloxacin, bleomycin) stress. Stallings et al. also showed that Arr interacts with L11 ribosomal protein (RplK). Interestingly, the function of L11 in modulating the translational GTPase and signalling synthesis of (p)ppGpp by RelA is a well-studied phenomenon.
In our current study (Agrawal et al., 2018), we have shown that arr confers a growth fitness advantage to M. smegmatis and contributes to biofilm formation, and the specific interaction of Arr with a ribosomal protein, L11 result in (p)ppGpp (guanosine penta/tetra phosphate, also known as “alarmone”) production in cells. Our study, for the first time, reports association of (p)ppGpp synthesis activity with the antibiotic resistance gene, arr, independent of any antibiotic treatment. The study has unravelled a novel role of antibiotic resistance gene arr in maintaining cellular homeostasis under environmental stress conditions. We may add that bacteria, when exposed to various adverse growth conditions, are known to respond by synthesis of (p)ppGpp. In these cases, the (p)ppGpp synthesis is primarily mediated by a protein called RelA. However, the role of Arr in (p)ppGpp production is an unusual one (Fig. 1).
Priyanka Agrawal, Umesh Varshney
Department of Microbiology and Cell Biology, Indian Institute of Science, Bangalore, India
Publication
Species-Specific Interactions of Arr with RplK Mediate Stringent Response in Bacteria.
Agrawal P, Varada R, Sah S, Bhattacharyya S, and Varshney U
J Bacteriol. 2018 Feb 23












Leave a Reply
You must be logged in to post a comment.