Are disinfection processes an effective barrier in controlling the spread of antibiotic resistance in the environment?
According to the report on global surveillance of antimicrobial resistance published in 2014 by the World Health Organization (WHO), the world is heading towards a post-antibiotic era, in which common infections and minor injuries may once again kill. Moreover, a team of scientist from Cambridge (UK) estimated that, depending on the scenario evaluated, failing to tackle antibiotic resistance (AR) threat will mean that the world population by 2050 may decrease by 11 million (if resistance rates will be successfully kept at a relatively low rate) and 444 million (it reflects a scenario for a world with no effective antimicrobial drugs) than it would otherwise be in the absence of AR. To successfully tackle AR threat not only clinical issue but also environmental one should be addressed.
The release of antibiotics into the environment through treated and untreated wastewater disposal contributes to the selection and development of antibiotic resistant bacteria (ARB) and antibiotic resistance genes (ARGs). ARB and ARGs (e.g., β-lactams (blaTEM), vancomycin (vanA) and tetracycline resistance genes) have been detected both in hospital effluents and in raw and treated effluent of urban wastewater treatment plants and consequently in surface water where these effluents are disposed of (Fig. 1). Accordingly, conventional disinfection processes (e.g., chlorination and UV radiation) may be not effective in controlling the release of antibiotic resistant mobile genetic elements in aquatic environments and they can promote horizontal gene transfer. Therefore alternative/new disinfection processes should be investigated to evaluate if they can minimize the risk of AR spread.
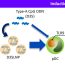
In this study an oxidant/disinfectant (hydrogen peroxide, H2O2) was coupled with UV radiation (UV/H2O2) to evaluate the effect on AR transfer through (i) the inactivation of an antibiotic resistant Escherichia coli (E. Coli) strain and (ii) the variation of selected ARGs (namely, blaTEM, qnrS and tetW genes). In spite of the complete inactivation of E. coli took place within 240 min of treatment by UV/H2O2 process (Fig. 2), no removal was reached for blaTEM gene (while the occurrence of qnrS and tetW genes was not unequivocally proven).

Fig. 2. Inactivation of antibiotic resistant Escherichia coli strain during disinfection by UV/H2O2 process.
Possibly, the disinfection process destroyed bacterial cells and promoted the release of ARGs in the water; therefore, ARGs may survive as dissociative DNA even if their bacterial hosts have been inactivated. According to the results achieved, UV/H2O2 process may be not effective in controlling the release of ARGs under realistic operating conditions. Consequently, if this process is used as advanced treatment of urban wastewater, ARGs unaffected by the process may be released into aquatic ecosystems where the effluent is discharged. Moreover, if the effluent is reused in agricultural practices, ARGs may be transferred to irrigated crops and soil. The pending question is if the released mobile genetic elements can still transfer antibiotic resistance to microorganisms in the receiving environment.
Luigi Rizzo
University of Salerno, Department of Civil Engineering, Fisciano (SA), Italy
Publication
β-lactams resistance gene quantification in an antibiotic resistant Escherichia coli water suspension treated by advanced oxidation with UV/H2O2.
Ferro G, Guarino F, Cicatelli A, Rizzo L
J Hazard Mater. 2016 Mar 7












Leave a Reply
You must be logged in to post a comment.