Deep Molecular Profiling to understand the biology of Alzheimer disease
Alzheimer disease (AD) is a complex, polygenic disease with genetic, cellular, pathologic, and clinical heterogeneity. Advances in high-throughput sequencing and omics technologies have accelerated the drive toward personalized medicine. Human bio-specimens using these high-throughput platforms have emerged as a critical resource for basic and translational research in neurodegenerative diseases because they can directly measure multiple layers of molecular data, from which targets for therapy, detection, and prevention are identified. A collection of human phenotypic data and their integration with “omics” data (genomic, proteomic, transcriptomic, epigenomic, lipidomics, and metabolomic, among others) provides an extraordinary opportunity for discovery of novel biomarkers, genes and pathways implicated in diseases and new therapeutic targets.
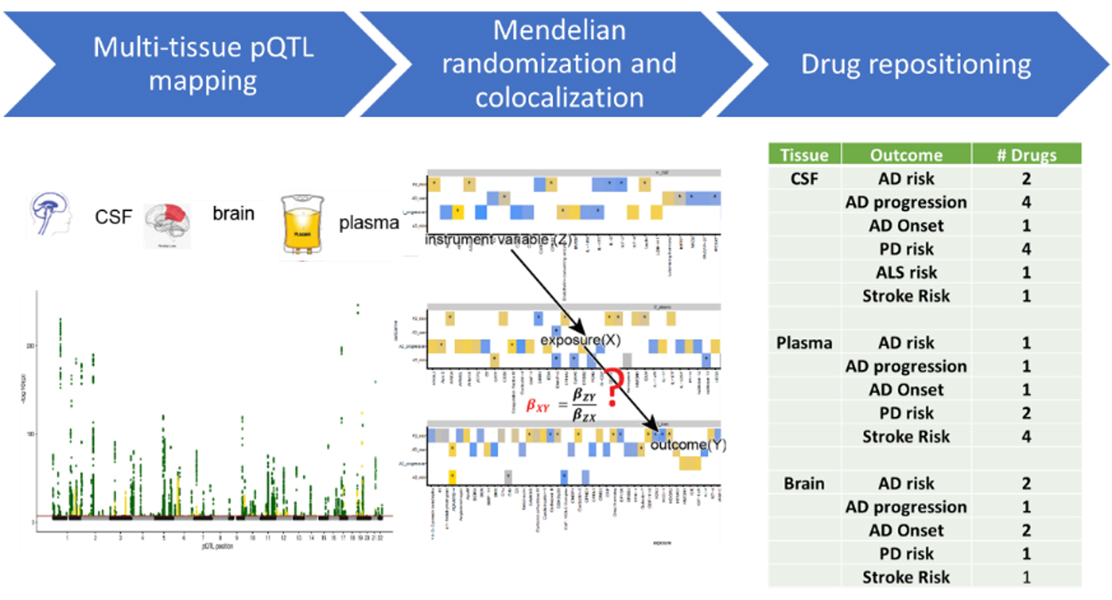
Fig. 1. Overall approach to identify casual genes and omic analytes and druggable targets. We combined pQTLs with Mendelian randomization and colocalization to identify proteins implicated on AD, PD, Frontotemporal dementia (FTD), amyotrophic lateral sclerosis (ALS) and stroke risk. We then examined if there are known FDA compounds that target those potential causal proteins.
Integrating genetic and multi-omic data provides a unique opportunity to identify new mechanisms that lead to understanding the biology of disease. Published eQTL studies have been instrumental in mapping the functional variants of GWAS loci. We and others demonstrate that other omic (including proteomic, methylation or metabolomics) data provide additional QTL information beyond eQTL. In this study, We quantified thousands of proteins brain, CSF and plasma from a large and well-characterized cohort. We identified 274, 127 and 32 pQTL for CSF, plasma and brain, respectively (Figure 1). We used AddNeuroMed, INTERVAL, KORA, and SCALLOP to replicate more than 98% of the pQTL. We demonstrated that cis-pQTL are more likely to be shared across tissues but trans-pQTL are tissue-specific. We demonstrated that 40-60% of the pQTL were no other type of QTL and we were able to map additional genes for AD and PD. By combining pQTL with Mendelian randomization (MR) and colocalization analyses, we were able to identify one CSF, 13 plasma, and six brain proteins that were likely causal for AD risk. Among them, plasma CD33 was a risk factor for AD and a drug target for other diseases, such as prostate cancer. For PD risk, 13 CSF, 12 plasma, and 23 brain proteins were likely in causal pathways, among which plasma IDUA was as a risk locus for PD and as a drug target for chondroitin sulfate, reported to treat osteoarthritis. We also identified several druggable targets and FDA compounds that could be repositioned for AD risk, progression, PD risk and other complex traits (Fig. 2)

Fig. 2. LRRK2 locus is a genetic modifier of Parkinson-associated genes (GRN, CSTB and GPNMB). A. PheWAS plot from the pQTL browser (Aim 3) for the common variant in LRRK2. B. Circos plot of the LRRK2 locus identifies the proteins regulated by LRRK2 variants
These results highlight the need for implementing additional functional genomic approaches beyond gene expression to understand the biology of disease and identify novel biomarkers and potential drug targets to improve personalized medicine.
Carlos Cruchaga
NeuroGenomics and Informatics Center, Washington University School of Medicine, St. Louis, MO, USA
Hope Center for Neurologic Diseases, Washington University in St. Louis, St. Louis, MO, USA
Department of Neurology, Washington University School of Medicine, St. Louis, MO, USA
Publication
Genomic atlas of the proteome from brain, CSF and plasma prioritizes proteins implicated in neurological disorders
Chengran Yang, Fabiana H G Farias, Laura Ibanez, Adam Suhy, Brooke Sadler, Maria Victoria Fernandez, Fengxian Wang, Joseph L Bradley, Brett Eiffert, John P Budde, Zeran Li, Umber Dube, Yun Ju Sung, , John C Morris, Anne M Fagan, Richard J Perrin, Herve Rhinn, Carlos Cruchaga
Nat Neurosci. 2021 Sep













Leave a Reply
You must be logged in to post a comment.