Determination of regions involved in amyloid fibril formation for Aβ(1-40) peptide
Mechanisms of amyloid formation of Aβ peptide are studied intensively. Determination of regions responsible for aggregation of the peptide and formation of amyloid fibrils is one of the primary aims in studying the formation of such structures. Knowledge of these regions will facilitate the choice of compounds interacting with a given region and thus preventing its aggregation. Therefore, in this study our task was to find amyloidogenic regions in amyloid fibrils formed by Aβ(1-40) peptide and to compare the data with the theoretical prediction. The FoldAmyloid method (http://bioinfo.protres.ru/fold-amyloid/) developed specially for this purpose was used to predict amyloidogenic regions of the Aβ(1-40) peptide.

Fig. 1. Sets of peptides accumulated after hydrolysis of Abeta(1-40) with a mixture of proteases (trypsin, chymotrypsin, proteinase K). The regions of Abeta(1-40) most protected from proteolysis in the amyloid structure are presented.
As known, the core of an amyloid fibril is resistant to the action of proteases. Therefore, it is possible to suggest that regions of the polypeptide chain that are involved in the core of the amyloid structure will be protected from the action of proteases, while the other parts of the polypeptide chain will be cleaved by proteases. Fibrils formed by this peptide were treated with a mixture of proteases consisting of trypsin, chymotrypsin, and proteinase K. When proteolysis was terminated, the mixture was centrifuged. The cleaved peptides (not included in the core of the amyloid fibril) remained in the supernatant, whereas amyloid structures were precipitated. The precipitate was washed, dried in a vacuum concentrator, and dissolved in formic acid. The sample was then layered on a reversed-phase column and eluted with an acetonitrile gradient. A mass spectrometer was used as the detector. The eluted peptides were ionized in the mass spectrometer, after which ions with the same mass were isolated and fragmented in an automatic mode using a specific algorithm. As a result, sets of masses of ions and their fragmentation spectra were obtained. After the analysis of the mass spectra and their fragments, the PEAKS Studio program was used to determine amino acid sequences of Aβ peptide fragments precipitated in the complex of amyloid fibrils.
Figure 1 shows peptides accumulated after hydrolysis of Aβ(1–40). It is seen that peptides of the C-terminal region in the Ab peptide are accumulated. In all C-terminal peptides, the region from amino acid residues 28 to 40 is resistant to the action of proteases. In addition, peptides are accumulated where the region from amino acid residues 16 to 22 also has increased resistance to the action of the proteases. In this case, control monomer (unassociated) structures of Aβ(1–40) are hydrolyzed completely into very short di-, tri-, and tetrapeptides.

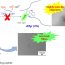
Fig. 2. Amyloidogenic regions predicted by the FoldAmyloid program are underlined by green color (http://bioinfo.protres.ru/fold-amyloid/).
Thus, using mass-spectrometry we determined regions of the chain that are protected from the action of proteases in amyloid fibrils formed by the Aβ(1–40) peptide. We demonstrated that for the synthetic Aβ(1–40) peptide, amino acid regions 16-22 and 28-40 are protected from the action of proteases upon formation of the amyloid structure. These data are in agreement with the amyloidogenic regions (amino acid residues 16-21 and 32-36) in the sequence of the Aβ(1–40) peptide predicted theoretically (Fig. 2).
Alexey K. Surin, Oxana V. Galzitskaya
Institute of Protein Research, Russian Academy of sciences
Publication
Determination of Regions Involved in Amyloid Fibril Formation for Aβ(1-40) Peptide.
Surin AK, Grigorashvili EI, Suvorina MY, Selivanova OM, Galzitskaya OV
Biochemistry (Mosc). 2016 Jul












Leave a Reply
You must be logged in to post a comment.