Determination of the size of folding nuclei of fibrils formed from Aβ(1-40) peptide
One of the most important models for studying amyloid fibril formation is the Aβ peptide, aggregates of which form plaques in the course of Alzheimer`s disease. Formation of Aβ aggregates goes through the nucleation mechanism with exponential acceleration of aggregation after seeds are formed. Though about 22000 papers (from the PubMed database) devoted to studying amyloid formation of Aβ peptide have been published, there is no consensus on the details of the scheme of aggregates formation. Previously we analyzed several models describing scenarios of amyloid fibril formation and gave equations, which connect experimentally measurable characteristic times with nuclei sizes and scenarios. As a direct consequence of equations, a scheme for analysis of amyloid kinetics was proposed. In this paper we have tried to apply such analysis and compared the obtained results to the electron microscopy (EM) images.

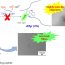
FIg. 1. Electron micrographs of a recombinant Aβ(1-40) incubated at 25 ° (50 mM Tris_HCl, pH 7,5): (a) after 55 h incubation; (b) gallery of separate annular oligomers (probable protofibril seeds) detected in the sample from 8 till 27 h incubation.
Formation of fibrils was traced by ThT fluorescence. Almost all kinetics had a lag-phase comparable to the aggregation time, and the analysis has shown the nucleus size to be around one monomer. This is a tricky situation since the nucleus size of one monomer may refer to several scenarios. One of the possible scenarios is the formation of some non-amyloid aggregates inside of which some rearrangements occur and seeds (the minimum size of stable state) are formed afterwards. It should be noted that according to the model, such “monomeric” seeds are building blocks for amyloid aggregates too. We obtained the EM images which support this scenario – thus, before formation of amyloid-like aggregates amorphous aggregates are present and then, after some time, ring-shaped oligomers are formed along with amyloid fibrils (Fig. 1). The aggregation process finishes when all non-fibril aggregates disappear. It is notable that though our data on the nucleus size do not coincide with our earlier estimations this could be addressed to the complex process of amyloid formation with high polymorphism of Aβ amyloid aggregates.

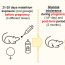
Fig. 2. Schematic representation of a possible mechanism of the formation of fibrils by Aβ(1-40) peptide: (a) destabilized monomer; (b) the fibril nucleus (primary oligomer, the basis for the formation of the annular oligomer); (c) the ring-like oligomer (protofibril, seed) composed of three primary oligomers is the basic building block for formation of fibrils; (d) the fibril build up from ring-like oligomers (protofibrils). The arrows show individual primary oligomers or the protofibril core.
The difficulty is that even during the lag phase one cannot state ex cathedra that the formation of fibrils starts from the monomeric form of Aβ(1-40). The amyloidogenesis process necessarily starts from destabilization of the monomer molecule. Such monomers form the nucleus of protofibrils. In the study of the kinetics of the formation of fibrils during the lag phase the estimated nucleus size will correspond to the primary oligomers containing several peptide molecules (3-6 molecules). Such oligomer is the basis for the formation of a larger annular oligomer / protofibrils (seed). In turn, the annular oligomers or protofibrils consisting of several primary oligomers are the basic building block for the formation of mature fibrils (Fig. 2). These annular mature fibrils and oligomers composed of ring-like oligomers were observed by the method of EM (Fig. 1). While at the beginning of the reaction of formation of fibrils ring-like oligomers can be detected (Fig. 2c), during the elongation of the incubation time only mature fibrils of Aβ(1-40) are left (Fig. 1a). It should be noted that the described mechanism of formation of fibrils from annular oligomers is presented for the first time. In addition, our calculations and the experimental data are quite consistent with each other.
Nikita V. Dovidchenko, Olga M. Selivanova, Oxana V. Galzitskaya
Institute of Protein Research, Russian Academy of sciences, Russia
Publications
Determination of Size of Folding Nuclei of Fibrils Formed from Recombinant Aβ(1-40) Peptide.
Grigorashvili EI, Selivanova OM, Dovidchenko NV, Dzhus UF, Mikhailina AO, Suvorina MY, Marchenkov VV, Surin AK, Galzitskaya OV
Biochemistry (Mosc). 2016 May
How to determine the size of folding nuclei of protofibrils from the concentration dependence of the rate and lag-time of aggregation. I. Modeling the amyloid protofibril formation.
Dovidchenko NV, Finkelstein AV, Galzitskaya OV
J Phys Chem B. 2014 Feb 6
The Mechanism Underlying Amyloid Polymorphism is Opened for Alzheimer’s Disease Amyloid-β Peptide.
Selivanova OM, Surin AK, Marchenkov VV, Dzhus UF, Grigorashvili EI, Suvorina MY, Glyakina AV, Dovidchenko NV, Galzitskaya OV
J Alzheimers Dis. 2016 Sep 6
One of the possible mechanisms of amyloid fibrils formation based on the sizes of primary and secondary folding nuclei of Aβ40 and Aβ42.
Dovidchenko NV, Glyakina AV, Selivanova OM, Grigorashvili EI, Suvorina MY, Dzhus UF, Mikhailina AO, Shiliaev NG, Marchenkov VV, Surin AK, Galzitskaya OV
J Struct Biol. 2016 Jun











Leave a Reply
You must be logged in to post a comment.