Enrichment design accommodating the biomarker misclassification
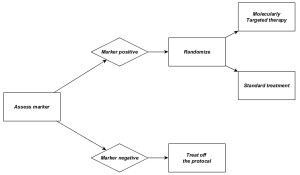
Increased understanding of the mechanistic heterogeneity of diseases has brought us to the era of molecularly targeted therapies (MTAs) in clinical oncology. The MTAs block the growth of cancer cells by identifying and attacking specific biomarker pathways that are deregulated in only a subset of patients and can only be expected to effectively treat a certain subset of patients (marker-positive). Therefore, to definitively evaluate the treatment effect of the MTAs, it is fair and sensible to restrict the trial enrollment to the marker-positive subgroup of patients whose tumors express the targeted biomarker. Based on this argument, the enrichment design was developed. As depicted in Figure 1, under the enrichment design, patients are classified into marker-positive and marker-negative subgroups based on their biomarker statuses, and only the marker-positive patients are randomized to receive different treatments including the MTA and standard therapy.
One practical obstacle to implement the enrichment design is that the precise measure of the biomarker status is often prohibitively expensive or logically infeasible for at least a subset of patients. Although a cheap and easy-to-obtain surrogate marker is often feasible for all the patients and can be used to predict the true biomarker status, such surrogate marker is often prone to the biomarker misclassification. The consequence of the biomarker misclassification is typically serious. From the view of trial implementation, due to the biomarker misclassification, the enrichment trial can include (exclude) marker-negative (positive) patients by mistake, which will undermine the ethics of the trial. From the view of data analysis, the misclassification causes bias and distorts the inference result. Therefore, appropriate accommodation is necessary and is in urgent demand.
We proposed a two-stage design to address this issue. In particular, we have developed an optimal rule to minimize the misclassification rate. Based on the optimal rule, we have developed the two-stage design. At the first stage, we measure both the precise and surrogate marker information and then equally randomize patients in this stage to receive different treatments. At the second stage, we only measure the surrogate marker and then use the optimal rule to make the marker classification. The patients classified as “marker-positive” are then enrolled into the trial and further randomized to different treatment groups whereas the patients classified as “marker-negative” are treated off the protocol. Hence, the optimal rule guarantees that the second stage patients have the minimal chance to make a marker classification error.
In addition to adapting the trial implementation, we also studied how to adjust the data analysis correspondingly. But just before that, we quantify the biases caused by the biomarker misclassification. Our derivation shows that the bias always exists regardless of the effectiveness of the MTA. Therefore, the biomarker misclassification can distort the conclusion of the enrichment trial, no matter whether the MTA is promising or not. To correct for the bias, we have developed a likelihood ratio test. To apply the likelihood ratio test, we need the maximum likelihood estimates (MLEs) for the response rates. However, due to the missing marker information in the second stage, these MLEs are difficult to derive directly. To solve this problem, we rely on the “Expectation-Maximization” algorithm, which is a well-known statistical method to handle missing data and derive the MLEs. After plugging in the MLEs based on the EM algorithm, we can determine the p-value for the test. Finally, this p-value is used to evaluate the MTAs under the enrichment trial with biomarker misclassification. We conducted comprehensive simulation studies to investigate the operating characteristics of the two-stage design and the results confirmed the desirable performance of the proposed design.
Yong zang1, Beibei Guo2
1Department of Mathematical Sciences, Florida Atlantic University, Boca Raton, FL, USA
2Department of Experimental Statistics, Louisiana State University, Baton Rouge, LA, USA
Publication
Optimal two-stage enrichment design correcting for biomarker misclassification.
Zang Y, Guo B
Stat Methods Med Res. 2015 Nov 26











Leave a Reply
You must be logged in to post a comment.