Fast and simple glyphosate detection using a functionalized surface
Glyphosate is the most widely used herbicide worldwide. It is used since the 1970s to dispose of weeds to increase yield. Since some years its use is under great debate because of its potentially cancerogenic effect and negative influence on biodiversity. Therefore, there is a high interest to get information about the presence of glyphosate in water, food, environmental probes etc. However, glyphosate detection is challenging due to its physicochemical properties like small size and high polarity. Usually, glyphosate is detected via chromatographic techniques which are cost and labor intensive and need qualified personnel. To overcome this problem, we aimed at developing a fast and simple assay that is widely usable by exploiting a completely new approach. Instead of detecting glyphosate itself or using antibodies, like in an ELISA-assay, we made use of the glyphosate target protein, the 5´-Enolpyruvylshikimate-3-phosphate synthase (EPSPS). The EPSPS is part of the shikimate pathway which is responsible for the formation of aromatic amino acids in plants and microorganisms. If glyphosate is present in the surrounding water or soil, the plant will starve because of the inhibition of EPSPS and the resulting lack of aromatic amino acids.
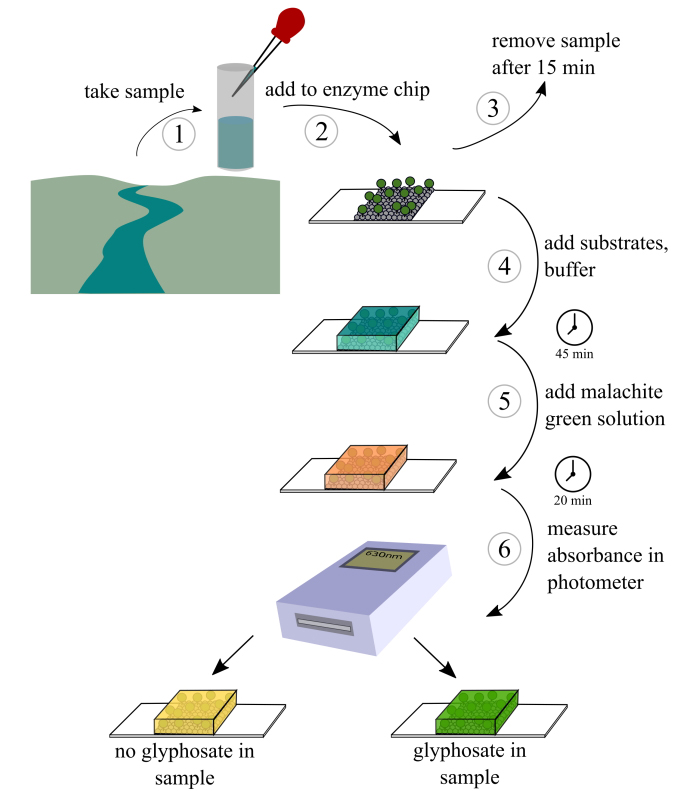
Using this enzyme, we wanted to provide an assay in a chip-format to make it easy to handle. We immobilized the EPSPS on a chip-surface using hydrophobin proteins from funghi. Hydrophobins are proteins with great properties. They are able to self-assemble at surfaces in a very stable and ordered manner. A fusion-protein of the EPSPS from the gut bacterium Escherichia coli (EcEPSPS) and the hydrophobin Ccg2 from Neurospora crassa was expressed and purified and used for the functionalization of plastic or glass surfaces which were used for the new assay. The assay is based on the reaction the EPSPS triggers. As substrates it uses shikimate-3-phosphate (S3P) and phosphoenolpyruvate (PEP) that are converted into enolpyruvylshikimate-3-phosphate and inorganic phosphate (Pi). In presence of glyphosate the enzyme is inhibited and the Pi level is lower compared to a control without glyphosate. The Pi is detectable using an easy colorimetric assay, the malachite green assay. With this laboratory setup, we were able to detect glyphosate in the nanomolar range (detection limit 50 nm). Furthermore, our assay showed great specificity for glyphosate. We tested different related and unrelated substances to glyphosate with our assay and documented, that they showed no cross reactivity. This is due to the high specificity of the glyphosate-EPSPS reaction. To our knowledge there is no other pesticide that specifically targets the EPSPS. This is an advantage compared to some other assays which showed cross-reactions with e.g. glyphosates first degradation product. However, indirect glyphosate detection via the malachite green assay is challenging as inorganic phosphates are ubiquitous in the environment. Therefore, we tested a preincubation method (compare to Fig. 1). We first added a phosphate and glyphosate containing solution to our surface and incubated for 15 minutes. During this incubation the glyphosate interacts specifically with the immobilized enzyme. Afterwards we replaced the solution with the assay buffer and performed the malachite green assay. We found out, that glyphosate inhibition was well measurable and the phosphate in the analyte solution was hardly interfering with our assay. With this method of operation, we were able to show the detection of glyphosate using its target enzyme, combined with an easy colorimetric assay.
Julia Döring, Kai Ostermann
Institute of Genetics, Technische Universität Dresden, Zellescher Weg 20b, 01217 Dresden, Germany
Publication
Surface Functionalization by Hydrophobin-EPSPS Fusion Protein Allows for the Fast and Simple Detection of Glyphosate
Julia Döring, David Rettke, Gerhard Rödel, Tilo Pompe, Kai Ostermann
Biosensors (Basel). 2019 Aug 29












Leave a Reply
You must be logged in to post a comment.