Frac-seq uncovers widespread RNA dysregulation in severe asthma
Asthma is a common chronic disease consisting of constriction and inflammation of the airways as well as structural changes known as remodelling (Fig. 1). 5-10% of patients have severe asthma, whereby symptoms persist despite regular and correct use of medication, including high dose corticosteroids. The causes underlying the lack of disease control remain poorly understood, highlighting that severe asthma is a critical unmet need.
The central dogma in biology states that DNA is transcribed into messenger RNA (mRNA), which is translated into proteins, the main cellular effectors. However, mRNA abundance does not always correlate with protein levels. Moreover, there are many RNAs that do not code for protein (non-coding RNAs) and function as regulatory molecules. We now know that the disconnection between mRNA and protein levels is mainly due to post-transcriptional regulation, carried out by microRNAs (small non-coding RNAs that inhibit gene expression) and RNA binding proteins. To date, most studies in severe asthma have centred in determining mRNA differences and inferring changes in specified proteins, without acknowledging posttranscriptional processes. To investigate posttranscriptional regulation, we employ Frac-seq (subcellular fractionation and RNA-sequencing). Frac-seq allows isolation and characterisation of mRNAs bound to polyribosomes (i.e. more than one ribosome, the translational machinery of the cell). This enables genome-wide comparison of total (transcribed) and polyribosome bound (undergoing translation) mRNA levels.
Bronchial epithelial cells (inner lining of the airways) are at the interface between the external environment and the tissue and play vital roles both as a barrier (structural) and as immune regulators. Investigating posttranscriptional dysregulation in severe asthma may offer new therapeutic targets and insight into disease. We thus implemented Frac-seq on bronchial epithelial cells from 8 severe asthma patients compared with 5 healthy controls and integrated those data with microRNA profiling employing small RNA-sequencing.
Our study revealed extensive posttranscriptional gene dysregulation in severe asthma, which is overlooked when only analysing total mRNA levels. mRNAs differentially bound to polyribosomes have little overlap with mRNAs differentially expressed at total levels between health and severe asthma and mapped to distinct biological pathways. For example, mRNAs differentially bound to polyribosomes showed enrichment in epithelium repair/remodeling pathways and absence of steroid signaling, the latter present only in differentially transcribed mRNAs. This may provide new insight into the lack of responsiveness to steroids seen in severe asthma.
Since microRNAs can modulate mRNA levels and/or translation, we investigated the genome-wide relationship between microRNAs and their target mRNAs. We detected 21 microRNAs differentially expressed in severe asthma in our Frac-seq cohort, which we cross-referenced with predicted targets present only in mRNAs differentially expressed in our Frac-seq datasets. We found little overlap between the targets in total vs polyribosome-bound mRNA, suggesting that dysregulated microRNAs have different effects on each subcellular compartment, showing a preference for mRNAs bound to polyribosomes.

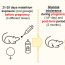
Fig. 2. Schematic of Frac-seq. Bronchial epithelial cells were isolated from 8 severe asthma patients and 5 healthy controls. Frac-seq was implemented. Small RNA-seq of the total RNA fraction was also performed. BECs: Bronchial epithelial cells.
To determine which of these 21 microRNAs play a more significant role biologically, we analysed the number of mRNA targets per microRNA. A network of only six microRNAs was predicted to undertake ~90% of microRNA targeting of differentially expressed mRNAs, with preferential targeting of polyribosome-bound mRNAs. Transfection of these microRNAs into bronchial epithelial cells from healthy donors mimicked severe asthma characteristics, e.g. rendering bronchial epithelial cells unresponsive to steroids.
Overall, our data show that there is widespread dysregulation of posttranscriptional processes in severe asthma and that a small network of microRNAs is responsible for a large proportion of these changes. Our work highlights the importance of integrating transcriptional and posttranscriptional data as well as assessing the relationship between master regulators and their targets, when investigating biological processes and disease.
Jennifer Rynne, Rocio T. Martinez-Nunez
Asthma UK Centre in Allergic Mechanisms of Asthma
School of Immunology & Microbial Sciences
Dept. Infectious Diseases, London, United Kingdom
Publication
Genome-Wide Posttranscriptional Dysregulation by MicroRNAs in Human Asthma as Revealed by Frac-seq.
Martinez-Nunez RT, Rupani H, Platé M, Niranjan M, Chambers RC, Howarth PH, Sanchez-Elsner T
J Immunol. 2018 Jul 1













Leave a Reply
You must be logged in to post a comment.