From parasitic microfungi towards the origins of Microsporidia
Microsporidia are unicellular eukaryotic parasites, infecting virtually all animals including humans. They are characterized by peculiar morphological and genetic features, which are the result of their extreme adaptation to the intracellular lifestyle. Interestingly, the infection apparatus in Microsporidia is very particular in that it is prearranged within the walled spore, and serves to inject the spore content into the host cell via a syringe-like polar filament. Once inside the host cell, the parasite multiplies by repeated fissions, and finally forms a new generation of walled spores. The Microsporidia also differ from the other eukaryotes, by lacking functional mitochondria (the energy-production central of the cell) stealing instead energy from the host cell, and by having compact genomes with high accelerated mutation rates and bacterial-like ribosomal RNA genes.

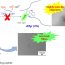
Fig. 1. The fungal kingdom comprises various lineages (phyla), of which the most basal are mainly aquatic and flagellated (zoosporic fungi), while more advanced forms have lost their flagellum and are predominantly terrestrial. It is still unclear whether Rozellomycota belongs to the Kingdom Fungi. However, present data strongly indicate that Microsporidia arose from this group, via intermediate forms represented by microsporidia-like organisms. The latter all share morphological similarities with primitive microsporidia, while molecular phylogenetic analyses show relatively close relationships with Microsporidia, although still not completely defined (lined box).
The evolutionary affinities of Microsporidia long time remained enigmatic, until recent genetic and genomic studies showed they are related to Rozella, a basal group of flagellated fungi (chytrids). Rozella species also are endoparasites, mainly of other water moulds, but their infection apparatus is not preformed inside the (flagellated) spore. By contrast, only after the parasite is attached to the host cell, it can generate an infection tube. In addition, although Rozella species have a mitochondrion, its genome (mt-genome) is greatly reduced, and Rozella steals the energy of the infected cell by the same mechanism used by the microsporidians i.e. by pumping ATP using a specific protein (ATP translocase) likely of bacterial origin.
Very recently new organisms have been identified, falling in the large diversity of the Rozella group (called Rozellomycota), that surprisingly resemble morphologically to microsporidia. Like microsporidians and in contrast to Rozella, these “microsporidia-like organisms”, i.e. Mitosporidium, Nucleophaga, and Paramicrosporidium, have non-flagellated spores with preformed (atypical) infection apparatus; however, the dramatic genetic modifications characterizing the microsporidians seem to be absent. None of these microsporidia-like organisms have visible mitochondria, however a mt-genome very similar to that of Rozella, is present in Mitosporidium, infecting the waterflea Daphnia (Crustacea) which was thus considered as an ancestral microsporidium. Nevertheless, phylogenetic reconstructions based on ribosomal RNA genes, showed that Mitosporidium is more closely related to a group of environmental clones than to microsporidians. Instead, Nucleophaga spp. which infect free-living amoebae result to be the closest relatives to Microsporidia.
The various phylogenetic analyses strongly indicate that these microsporidia-like organisms are to various extents affiliated to Microsporidia, but also to various environmental clones within the Rozellomycota. Furthermore they also seem to share some morphological details with a poorly studied group of putative primitive microsporidia. Overall, these results suggest that the recovered microsporidia-like organisms are potential intermediate forms, and that the peculiar morphology of Microsporidia has evolved early from these parasitic rozellomycotans.
In conclusion, while the protistan or early fungal nature of the Rozellomycota is a matter of debate, it is clear that the Microsporidia have emerged from inside this group of intracellular parasites. Future comparative genomic studies of the various intermediate forms will allow to better elucidate the transitional steps of this very interesting lineage of organisms.
Corsaro D
CHLAREAS Chlamydia Research Association Nancy, France
Publication
Molecular identification of Nucleophaga terricolae sp. nov. (Rozellomycota), and new insights on the origin of the Microsporidia.
Corsaro D, Michel R, Walochnik J, Venditti D, Müller KD, Hauröder B, Wylezich C
Parasitol Res. 2016 Aug












Leave a Reply
You must be logged in to post a comment.