Genetics of mice and men: NF1 patient-specific mouse models offer hope
Neurofibromatosis type 1 (NF1) is one of the most common inherited neurological disorders, affecting about 1 in 3,000 people throughout the world. The disorder is characterized by light brown skin spots (café-au-lait spots) and small benign growths (neurofibromas) on or under the skin. Occasionally, tumors develop in the brain, on cranial nerves, or near the spinal cord.
NF1 is caused due to changes (mutations) at specific places within a person’s genetic information. We all have a large amount of genetic information organized into smaller units known as genes. Genes provide the basic instructions and information that our cells need to perform their duties within our bodies.

Fig. 1. Impact on protein from healthy and mutated genes. Both missense and nonsense mutations are caused from a single change while the final outcome differs greatly.
In patients with NF1, the disorder develops as the result of mutations in a gene that codes for the protein neurofibromin. Neurofibromin acts as a tumor suppressor and helps keep cells from growing and dividing too quickly. In the absence of neurofibromin, certain signaling pathways in the cell are no longer held in check. This leads to abnormal growth and division of the cells, increasing their chance to develop to tumors. Manifestation of NF1 vastly differs from patient to patient, and no one person will have all of the possible symptoms of NF1. One reason for this is the unique genetic mutation(s) that affects NF1 patients.
We are interested in developing therapies that target specific NF1 genetic mutations. In order to do that we need to understand how the disorder is caused and animal models with these patient-specific mutations can help immensely. Therefore, we developed mice harboring two NF1 patient-specific mutations including a missense mutation (c.2542G>C; p.Gly848Arg) and a nonsense mutation (c.2041C>T; p.Arg681*). A missense mutation acts like a detour sign within a gene by causing a single change in the amino acid composition of the very large protein (Fig. 1B). Protein is still produced but it may or may not function properly. This seemingly minor change has the potential to interfere with neurofibromin function. A nonsense mutation is a single change that creates an early stop sign signal within a gene and does not allow the gene to code for a full-length protein (Fig. 1C). The results is the production of a shorter protein or in extreme situations no protein at all. The particular missense mutation that we studied is associated with the development of multiple spinal tumors, whereas the nonsense mutation is associated with café au laits, dermal and plexiform neurofibromas, and no spinal tumors in NF1 patients (Fig. 2A).

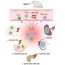
Fig. 2. Comparison between NF1 patients and mice with the same NF1 mutations. NF1 patient photos courtesy of Dr. Bruce Korf, UAB Neurofibromatosis Program. Disease Models & Mechanisms original article – http://dmm.biologists.org/content/9/7/759
From our experiments, we observe that the human nonsense and missense mutations have different effects on neurofibromin protein expression in the mouse and each reveals unique aspects of the NF1 disorder. The missense mutation fails to produce the disorder in the mouse as compared to the spinal tumors observed in NF1 patients with this mutation (Fig. 2B). However, when put into the appropriate genetic context to knockout the Nf1 gene in endothelial cells and Schwann cells of the peripheral nervous system, the nonsense mutation does cause spinal tumors that are not observed in NF1 patients with this mutation. These tumors compress the spinal cord and cause paralysis (Fig. 2B). Human and mouse are different organisms, and so these findings are not completely unexpected. The mouse is a great experimental tool to study human disease, and our observed differences are helping shed light on better understanding the NF1 disorder.
This nonsense NF1 mouse model will be valuable for preclinical testing of novel nonsense suppression therapies. These new therapies target in-frame nonsense mutations that lead to the disorder in individuals with nonsense mutation NF1.
Ashley N. Turner
Department of Genetics & UAB Neurofibromatosis Program,
University of Alabama at Birmingham,
Birmingham, Alabama, USA
Publication
Mice with missense and nonsense NF1 mutations display divergent phenotypes compared with human neurofibromatosis type I.
Li K, Turner AN, Chen M, Brosius SN, Schoeb TR, Messiaen LM, Bedwell DM, Zinn KR, Anastasaki C, Gutmann DH, Korf BR, Kesterson RA
Dis Model Mech. 2016 Jul 1












Leave a Reply
You must be logged in to post a comment.