Genome editing in non-dividing cells: new strategies are required
Although rare individually, genetic disorders collectively constitute a common health problem. As the cause of these diseases is a defective gene, gene therapy would be able to resolve all of these disorders. The most recent method of gene therapy is called gene editing. Gene editing is repairing the defect of the gene by using molecular scissors such as CRISPR/Cas9 (Fig. 1). CRISPR/Cas9, which is the most commonly used type of this system is, composed of two components Cas9 protein which can make a cleavage in DNA and a guide RNA that is a RNA molecule which binds to Cas9 and guide it towards a specific target sequence of 20 bases. CRISPR/Cas9 allows us to cut the human genome by generating DNA double-strand breaks (DSB) close to the mutation. Such DSB is sensed by two main cellular repair pathways, nonhomologous end-joining (NHEJ) and homology-directed repair (HDR).

Fig. 1. CRISPR/Cas9 system. Cas9 and sgRNA targeting a sequence protospacer, provided that a 3’ PAM sequence is present (NGG). The cutting site is shown with black scissor and located 3 bases upstream from the PAM sequence. (Nami et al., 2018).
While NHEJ is generally known to be an error-prone pathway causing insertions and deletions (indels) of DNA bases, which occurs independently of a repair template, HDR relies on the presence of a repair template to refill the correct bases and is considered an error-free pathway. Therefore, HDR has been extensively tested and used for precise gene editing. Given that HDR is restricted to the S and G2 phases of the cell cycle, only present in dividing cells, this approach might not be suitable for editing and repairing the genome in non-dividing cells. However many of monogenic disorders, cause disease, because the function of terminally differentiated non-dividing cells in different organs is impaired. Therefore, to reverse the defect caused by monogenic diseases, therapies that can correct the genetic defect in non-dividing cells are required. Recently, several novel genome-editing strategies, have been developed, which might make it possible to correct genetic mutations in non-dividing cells. These strategies can be categorized into two main groups based on their dependency on the repair (donor) template (Fig. 2). The donor dependent methods are mainly based on inserting a piece of DNA while donor independent strategies are based on deleting, making indels, changing a base or inverting the target sequence. Therefore, donor dependent strategies are mostly useful for reversing large-scale mutation such as big deletions while donor independent strategies are beneficial mostly for reversing gain of function mutations by disturbing the sequence of the mutated gene. It should be mentioned that none of these strategies allow for precise and scarless repair of mutations except for base editing. Base editing is a direct replacement of a single DNA base with the correct one without making a DSB and therefore not relying on HDR. Base editor (BE) is an engineered fusion enzyme consist of Cas9 and cytidine deaminase (CD) that enables a C-G to T-A conversion in an activity window that can be as narrow as 1-2 bases. Since the initial description, BE has been extensively improved and expanded.

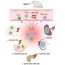
Fig. 2. Candidate gene editing approaches for non-dividing cells in vivo.
(a and b) Donor-independent approaches, (c and d) donor-dependent approaches. Boxes with dash outlines are potential approaches which insufficient experimental proof that they can edit genes in vivo in non-dividing cells exists (b and d). In each box approaches (in bold letters) and strategies are listed in the left column, targetable mutations for each approach are listed in the middle column, and schematic representation of each approach is shown in the right column.
(right column): an exon in pink color, and a red star indicate the mutated exon; dashed lines show splicing pattern; green triangles represent the manipulation site; Stop: stop codon; PA: poly A sequence.
(box ‘a’): the cryptic exon (crypt ex) is shown as an example.
(box ‘c’): pink exon2 with a jagged side represent a deletion.
(abbreviations used) BE: base editor, BE 3, BE4 and YEE BE3 are other BE developed from BE3, HITI: Homology-independent targeted Integration, HMEJ: Homology-mediated end Joining, ObLiGaRe: Obligate Ligation-Gated Recombination, PITCH: Precise integration into target Chromosome, recCas9: recombinase Cas9, Target-AID: a base editing system very similar to BE3 using a different cytidine deaminase. (Nami et al., 2018).
Another important base editing tool is adenine base editors (ABEs) contains a DNA adenosine deaminase instead of CD and enables a T-A to C-G transition. ABE is a major milestone in base editing considering that the target mutations (C-G to T-A ) account for half of the pathogenic known point mutations in human. One important limitation of both BE and ABE is that the point mutation should fall within the activity window in the target sequence. Additionally it is still not possible to reverse the Transversion (interchanges of purine for pyrimidine bases) mutations or other small-scale mutations such as indels using base editing. Therefore, development of other genome editing strategies for precise repair of small-scale mutations are required.
Fatemeharefeh Nami 1, Hossein Baharvand 2,3 , Catherine Verfaillie 1
1Department of Development and Regeneration, KU Leuven-Stamcelinstituut, Leuven, Belgium
2Department of Stem Cells and Developmental Biology, Cell Science Research Center, Royan Institute for Stem Cell Biology and Technology, ACECR, Tehran, Iran
3Department of Developmental Biology, University of Science and Culture, Tehran, Iran
Publication
Strategies for In Vivo Genome Editing in Nondividing Cells.
Nami F, Basiri M, Satarian L, Curtiss C, Baharvand H, Verfaillie C
Trends Biotechnol. 2018 Aug












Leave a Reply
You must be logged in to post a comment.