Human cells can count their chromosomes – but how do they do?
Human cells normally carry 46 chromosomes. There are 2 gonosomes and 22 pairs of autosomes, numbered from 1 through 22. In male there are as gonosomes an X- and a Y-chromosome; in female there are two X-chromosomes. It is well known that for regular function of female cells the majority of genes present on one of the two X-chromosomes needs to be blocked. This mechanism is called ‘X-chromosome-inactivation’. Recent research showed an essential step during early embryogenesis is, that fetal cells check if there is only one or if there is more than one X-chromosome in their nucleus. In other words, these cells are able to determine the number of X-chromosomes per cell. Or more precisely: these cells are able to count their own X-chromosomes. This is possible by physical interaction between all X-chromosomes present in a nucleus at a certain phase of the cell cycle; this phenomenon was given the romantic sounding name ‘chromosome-kissing’ (Fig. 1A). The gene which is involved in this chromosome-kissing-event is called “solute carrier family 16, member A2” or in short SLC16A2 gene.

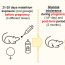
Fig. 1. A) The gene SLC16A2 is incolved in X-chromosome (X-chr.) counting.
B) Almost each human autosome carries a pseudogene/ gene highly homologous to SLC16A2 (SLC16A2-like). Thus, those genes could be involved in autosome counting.
For a functional and healthy human individual, besides correct X-chromosome numbers also modal chromosome numbers of all other human chromosomes (the 22 autosomes) must be correct. Interestingly, there are hints from clinical genetic diagnostics that also for other chromosomes besides X-chromosome, counting mechanisms must exist during early embryogenesis. The latter lead to correction of trisomies or monosomies of single chromosomes, either by degradation of a superfluous chromosome or by duplication of a single chromosome. These counting mechanisms work less efficiently than that involved in X-chromosome inactivation, however, they exist.
Recently a database search for sequences being homologous to the X-chromosome-specific SLC16A2 gene was performed for all human autosomes. It revealed that such homologies exist practically along all human chromosomes 1 to 22. Thus, the “SLC16A2 gene-family” may be a promising candidate for a better understanding how human cells determine their correct modal chromosome numbers. However, this would be just a starting point for future research, as yet it is completely unclear how the cells e.g. degrade a supernumerary chromosome or duplicate specifically a monosomic one.
Thomas Liehr
Jena University Hospital, Friedrich Schiller University, Institute of Human Genetics,
Kollegiengasse 10, D-07743 Jena, Germany
Publication
Thoughts about SLC16A2, TSIX and XIST gene like sites in the human genome and a potential role in cellular chromosome counting.
Rinčić M, Iourov IY, Liehr T
Mol Cytogenet. 2016 Aug 8












Leave a Reply
You must be logged in to post a comment.