Molecular origins of convergence and uniqueness of protein structures!
Protein folding, tagged as an NP hard problem is the process by which a protein sequence acquires its biologically functional native structure, with a high-fidelity rate. Proteins are essential components of all body tissues, muscles and act as important mediators of all cellular processes in the form of blood, hormones, antibodies, enzymes etc.
Proteins use a combination of 20 monomers (called amino acids) in certain allowed stoichiometries to create polymers with unique structures and this is maintained through the evolutionary course of time. The Protein data bank, (RCSB), a structural repository of biomolecules currently hosts ~135000 protein structures. Where is the unique structure of a protein coming from?

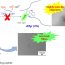
Fig. 1. (A): A self-avoiding random coil like polymer which is made up of n-mers that can freely rotate across any bond, (B) protein chains are highly organized tight structures which can oscillate only about a fixed region in the dihedral space, rather than excursing into myriads of conformational possibilities (PDB ID 1MBN, adopted from Protein Data Bank).
In early 1960s, Anfinsen hypothesized that the amino acid sequence dictates the unique three-dimensional structure of a protein. How does nature do this incredibly complicated task of weaving these unique structures from sequences routinely that too in millisecond-second time scales?
Proteins are characterized by two backbone degrees of freedom, the dihedral angles phi and psi (φ, ψ) around each Cα atom. Each dihedral can take a maximum value of 2π radians, i.e. the conformational space around each Cα is 4π2 radians, and for an ‘n’ residue protein, the total conformational volume is (4π2)n which tends to infinity with increasing n (i.e. divergent). Ramachandran and coworkers, discovered that most of the dihedral space is disallowed due to backbone atom clashes when analyzed at the monomer level. Introduction of a function, f (φ, ψ), for allowed fraction, converts the conformational volume to (4π2f)n. Is this volume divergent leading to innumerable conformations or convergent leading to a unique structure?
Mathematically, the value of 1/f should be greater than 4π2 to ensure convergence of protein structures. A systematic exploration of Ramachandran’s idea of disallowed regions due to steric clashes at the monomer, dimer, trimer, tetramer and pentamer levels from 43612 crystal structures resulted in f values of 0.26, 0.054, 0.024, 0.019 and 0.018 and 4π2f values of 10.3, 2.1 , 0.9, 0.8 and 0.7 respectively. While an increase in the values of n, leads to divergence in the former two cases, the latter three cases show shrinkage to zero as the chain grows. Thus the origin of uniqueness in protein structures is inherent in the sequence at the tripeptide level and beyond. Hence tripeptides and larger peptide segments have enough information to restrict the conformational volume to lead to the creation of unique structures.
A protein can thus accommodate locally rigid and flexible regions through fine-tuning the allowed fraction f by choosing appropriate trimers and multimers in their sequences.
Debarati DasGupta 1,2, B. Jayaram 1,2,3
1Department of Chemistry,
2Supercomputing Facility for Bioinformatics & Computational Biology &
3Kusuma School of Biological Sciences,
Indian Institute of Technology, Hauz Khas, New Delhi, India
Publication
Protein folding is a convergent problem!
Das Gupta D, Kaushik R, Jayaram B
Biochem Biophys Res Commun. 2016 Nov 25












Leave a Reply
You must be logged in to post a comment.