Short hydrogen bonds in proteins and their quantum mechanical nature
The assembly of linear polypeptide chains into functional three-dimensional protein architectures involves a unique force called hydrogen bonding. A typical hydrogen bond forms when the donor and acceptor groups approach each other with a nearly linear geometry and a heteroatom distance between 2.8 Å and 3.2 Å. Interestingly, the protein folds often allow amino acids to reside in much closer proximity and form hydrogen bonds with the heteroatom distances below 2.7 Å. We have statistically analyzed the Protein Data Bank, which contains over 153000 biomolecular structures, to identify the occurrence of these short hydrogen bonds (SHBs) and characterize their structural and chemical features. Here we focus on the top 1% highest quality structures in the database and identify that the O-H•••O, O-H•••N, N-H•••O and N-H•••N type SHBs and their networks are prevalent in proteins and protein-ligand complexes. They hold amino acids together and connect active-site amino acids and ligands (Fig. 1), and might contribute to essential biological functions. SHBs are particularly enriched in enzymes, the biological catalysts that accelerate chemical reactions in living organisms (Fig. 1). For example, they occur extensively in hydrolases and possibly assist the cleavage of chemical bonds using water molecules. They are also commonly found in oxidoreductases and potentially facilitate the efficient transfer of electrons between molecules.

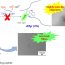
Fig. 1. Results from analyzing 1663 high-resolution crystal structures from the Protein Data Bank. (a) Distribution of 11814 SHBs in enzymes. Example structures of SHBs (b) formed between amino acids and (c) formed between an active-site amino acid and ligand in proteins.
While hydrogen bonds are often considered as classical dipole-dipole interactions, SHBs exhibit prominent covalent characters. Since hydrogen is the lightest element, nuclear quantum effects can also significantly alter their geometries and properties. We have conducted quantum chemistry calculations of 3665 SHBs formed from the side chains of amino acids and obtained the potential energy surfaces for moving the proton between the donor and acceptor atoms. The resulting surfaces can take the shape of a double well (Fig. 2a), single well with a shoulder (Fig. 2b) or single well (Fig. 2c). As a hydrogen bond shortens, it is more likely to have a single well potential with a low energy barrier and a shared proton. In many SHBs, the energy barrier is comparable to the zero-point energy of a typical O-H or N-H bond (~5 kcal/mol) and the proton becomes less confined around the donor atom. As such, nuclear quantum effects allow the proton to move towards the midpoint between the donor and acceptor atoms and further enhance the covalency of the hydrogen bond.

Fig. 2. Example potential energy surfaces for moving the proton in a SHB. In the plots, the proton is closer to the hydrogen bond donor or acceptor atom when its position is negative or positive, respectively. The horizontal line represents the zero-point energy of a typical O-H or N-H bond.
Our findings extend the knowledge of hydrogen bonds in biological macromolecules. We demonstrate that SHBs widely exist in proteins and have distinct quantum mechanical nature. As such, they do not follow the standard description of hydrogen bonds as classical electrostatic interactions. Our work will facilitate the incorporation of these close contacts in the experimental and computational refinement of protein structures and improve the modeling of the structures, interactions and functions of these biomolecules.
Shengmin Zhou 1, Lu Wang 2
1YDS Pharmatech, Inc., Albany, NY, USA
2Department of Chemistry and Chemical Biology, Institute for Quantitative Biomedicine, Rutgers University, Piscataway, USA
Publication
Unraveling the structural and chemical features of biological short hydrogen bonds
Shengmin Zhou, Lu Wang
Chem Sci. 2019 Jul 1












Leave a Reply
You must be logged in to post a comment.