The binding of mGluR4 targeting drugs could help the finding novel treatments for PD
One of the key features of drugs is an ability to find the corresponding target molecule (eg. protein) from the body. The target protein has unique structural sites that could be loaded with specific chemicals (drugs). The drug has to find its way to the target and bind to the specific site “hot spot” to be able to modulate biological effects in body. The development and optimization of target specific drugs is demanding, since the human body consists of thousands of different drug targets, which have potential to bind the drug and possibly cause unwanted side effects. Like in our study, simplified models of human body were developed and used to mimic the environment in order to evaluate, screen and estimate drug like properties of drug candidates in safer, more economical and less time consuming manner.
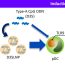
Our study presents a ligand binding assay, which was performed with cells that were specially engineered to overexpress our target protein, metabotropic glutamate receptor 4 (mGluR4). This model allows studying the formation of drug-receptor complex using radioactive drug molecule in a simplified and more regulated biological environment. The radioligand binding- assay (RIA) is a classical approach, where the drug-target complex is formed and the unbound radiolabeled reference drug is washed from the cell samples and the amount of bound drug could be measured.
In fact, binding assays are one of the first steps of drug discovery processes when new drug candidates are tested, evaluated and optimized. For example, Parkinson disease (PD) is classically treated with L-DOPA, which works in an early stage of the disease and after a few years the disease can progress to a resistant phase and the patients start to have additional symptoms despite of the ongoing treatment. Glutamate is a neurotransmitter and the mGluR4 is primarily expressed in the presynaptic site of the synapse and it plays an important role in the basal ganglia, which is one of the most essential areas of the brain linked to PD. The glutamate binding activates the mGluR4 and reduces transmission at synapses, which are overacting in PD. Glutamate binding moiety is found from many other proteins in humans, which makes it extremely hard to find specific drugs to the glutamate binding site in mGluR4. Fortunately positive allosteric modulators (PAMs) of mGluR4 have been developed. PAMs binds to distinct site of the mGluR4, but could potentiate and enhance the glutamate signaling. Unfortunately only little is known about the binding of mGluR4 PAMs. Ligand binding studies are common in early stage drug discovery for optimizing the selectivity, but also give the ability to quantify receptor densities when drugs are developed as imaging agents. Our studies revealed that the lead PAMs had co-operative binding with glutamate, which means that the binding of glutamate increases the binding of the investigated PAMs to mGluR4.
These kind of simple binding assays have major impact on selection of potential drugs and imaging agents and optimizing them. The characteristic co-operative factor of the binding of mGuR4 PAMs should be taken account when new drugs are tested and designed. Simple binding assays combined with other drug evaluation tests will further our understanding of the potential of mGluR4 modulation as a novel symptomatic and potentially disease-modifying treatment for diseases like PD. We are eagerly waiting to see the first mGluR4 imaging agents available for clinical applications to further visualize the mGluR4 actions in patients and hopefully clarify the mechanism of development and progression of multiple neurodegenerative diseases including PD. Ultimately our aim is to find cure for the patients that suffer from diseases that are untreatable at the moment.
Publication
Co-operative binding assay for the characterization of mGlu4 allosteric modulators.
Poutiainen P, Kil KE, Zhang Z, Kuruppu D, Tannous B, Brownell AL
Neuropharmacology. 2015 Oct












Leave a Reply
You must be logged in to post a comment.