Transcription modulation in Xanthomonas citri subsp. citri
Xanthomonas citri subsp. citri (X. citri) is a Gram-negative bacterium and the etiological agent of citrus canker, a severe disease that affects all the commercially important citrus varieties with worldwide distribution. The best method to control the spread of this plant pathogen, which happens by the combined action of wind and rain, is the eradication of symptomatic and asymptomatic trees in the orchards. However, a conjunction of political and economic circumstances led to a scenario in which eradication was abolished from the beginning of 2017 in the state of São Paulo, Brazil; the main orange producing area, and the largest orange juice producer in the world. Currently, control is achieved by the intensification of agricultural integrated management practices that require, among others, recurrent sprays of copper formulations, a potentially toxic metal that accumulates in the soil and water reservoirs.

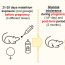
Fig. 1. Depletion of ParB-TAP in X. citri ΔparB. The bacterial citrus pathogen X. citri, deleted for the chromosome segregation gene parB and expressing ParB-TAP from the replicative vector pLAC2, was cultivated optimally from the starting O.D.600nm of ~0.1. After 6h of growth, the promoter inducer arabinose (0.05%), and the repressor glucose (2%) were added to the cultures (black vertical arrow in A). O.D.600nm measurements were carried out every 6h and used to construct the growth curves shown in (A). B) Depletion window of ParB-TAP. Cell samples were extracted from the curves in A at the time-points indicated, and processed for the immune-detection of ParB-TAP in western blot. The time 6h (lanes 1-2) shows the basal expression of ParB-TAP from the replicative vector pLAC2, before the addition of any sugar; note the absence of ParB-TAP at 13h and 19h of exposure to glucose; at these time points cells are already elongated and cultures may exhibit increased numbers of anucleated rods, which is suggestive of chromosome segregation defects (C). Lanes 3 and 6 in B are empty.
Our group has long been committed to develop alternative methods to control citrus canker, which are based on the use of more effective and necessarily less toxic/environmental friendly compounds able to disrupt vital cellular processes of X. citri. An important step to accomplish this task is the possibility of performing genetic and biochemical characterization of proteins that could be potential targets to be disrupted. Unfortunately, finding targets has not been an easy job in this plant pathogen, especially when the gene/protein under investigation is essential for living. An alternative to circumvent this problem in several organisms has been the possibility of modulating transcription or translation without gene removal. In this work, we report on the construction and characterization of the pLAL/pLAC series of protein expression vectors intended for protein depletion studies in X. citri. Our vectors carry the arabinose promoter derived from the pBAD series largely explored in Escherichia coli. The X. citri protein expression system enabled controllable protein expression in this phytopathogen, displaying a remarkably fine capability for transcription regulation with little promoter leakage (if using the replicative vector pLAC) or no leakage at all (when using pLAL derivatives that function integrated as a single copy into the bacterial chromosome) (Fig. 1). To validate them, pLAL vectors were used as complementation tools for the clean deletion of parB in X. citri, a widespread and conserved gene involved in the essential process of bacterial chromosome segregation. Following parB knockout, we used pLAL-parB to ectopically express ParB within X. citri. The effects caused by the imbalance of ParB in X. citri were investigated by modulating ParB expression. Both, ParB over-production or depletion led to chromosome segregation defects, without interfering with the ability of the X. citri DparB mutant strain to colonize the host citrus and cause disease. Finally, the arabinose expression vectors described here are valuable tools for protein expression, gene complementation, protein depletion, and especially, to assist in the deletion of essential genes in X. citri and probably related phytopathogens.
Henrique Ferreira
Depto. de Bioquimica e Microbiologia, Instituto de Biociencias de Rio Claro,
Universidade Estadual Paulista, Avenida 24-A 1515, Bela Vista, Rio Claro, Brazil
Publication
Protein depletion using the arabinose promoter in Xanthomonas citri subsp. citri.
Lacerda LA, Cavalca LB, Martins PMM, Govone JS, Bacci M Jr, Ferreira H
Plasmid. 2017 Mar












Leave a Reply
You must be logged in to post a comment.