What about the real costs of next generation sequencing (NGS) in human genetic diagnostics?
Human genetics diagnostics (HGD), which always has to go together with human genetic counselling, is a rapidly developing field. HGD may become necessary in case of familial cancer syndromes, infertility, prenatal abnormal sonography or a child born with dysmorphic signs, developmental delay and/or mental impairment.

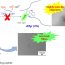
Fig. 1. Costs in connection with next generation sequencing (NGS) shown schematically – those which are normally considered and discussed (in black), those which are normally not considered (in green).
While at the beginning of routine HGD, in the 1970s, banding cytogenetics was the only available tool for chromosome and DNA analyses, nowadays a whole battery of approaches is accessible. All of these newer techniques went through a process of application (1) only in ‘pure research’, then (2) in ‘research and practical cases in parallel’, to (3) an ‘established procedure in routine’. Approaches like polymerase chain reaction (PCR) based applications, Sanger sequencing or molecular cytogenetics and molecular karyotyping finished these procedures already latest 5-10 years ago. The most recent technical development, which at present is quickly entering HGD, is so-called next generation sequencing (NGS).
NGS can be based on different, highly sophisticated principles finally leading to the deciphering of a human sequence. It is a fascinating approach, by which, compared to Sanger sequencing, incredible quickly sequencing results can be obtained. In terms of maturity and stability of NGS, it is an advanced technology being really an ‘established procedure in routine’ (i.e. arrived at step 3 from above).
Still, data interpretation is not that mature yet, as many variants from the ‘standard genome’ can neither be assigned as “benign variant” nor as “disease causing variant”, but have to be classified as “variants of unclear significance”. Even worse: during the last years many initially as “variants of unclear significance” classified sequence deviations had to be re-classified as “benign variant” or even as “disease causing variant”; also “disease causing variants” were downgraded to “variants of unclear significance” or even to “benign variant”. Thus, concering result interpretation NGS is still on “stage 2” as defined above, i.e. ‘research and practical cases in parallel’ performed. The latter is nothing new in HGD, as in the beginning of banding cytogenetics similar things were observed with this, at that time, new approach. That is simply the fact owed that it cannot be assumed before patients not to offer them the new possibilities NGS in case there is a good chance that it may help to solve his / her problem.
However, there are two closely linked argumentations in connection with NGS and its application in HGD, which are really puzzling: the statement (i) that NGS is a cost efficient approach and will become cheaper and cheaper with time, and thus (ii) should replace all other routine approaches in HGD. Funnily enough both statements were expressed about 10 years ago just substituting the word ‘NGS’ by ‘molecular karyotyping’; none of both suggestions for molecular karyotyping became true by now.
Additionally, one must admit that running NGS in HGD needs not only the (maybe in future a little bit, but not much) cheaper consumables (like cartridges, flow cells and chemicals), but there are other, in main stream publications always not considered costs of NGS (Fig. 1):
- First, to be allowed to do accredited HGD (at least in Europe and North-America) each NGS machine has to undergo yearly maintenance by the providing company, leading to costs in the range of 10.000 to 50.000 Euro.
- Second one has to participate in quality control studies (costs of 1.000 to several thousand Euros per anno).
- Third, NGS data has to be evaluated by the support of software programs, which may contain yearly charges in the range of 10.000s of Euro.
- Finally, specialists (optimally bioinformaticians) are needed, for developing optimal data evaluation pipelines; biologists / MDs for clinical relevant evaluation of the data are the second group of professionals to be hired and paid (Fig. 1).
According to the question, evaluation of one case done by NGS needs in simple cases at least 2 hours; in exceptional cases one person can work on the result for several days to weeks; the latter is, among others, due to the necessity to classify hundreds of sequence variants per case as “benign”, “disease causing” or “variants of unclear significance”, before being able to come to a final conclusion about the meaning of the obtained results.
Overall, NGS is a new tool already successfully applied in HGD in multiple cases. In the years to come it will complete the tools available for good patient care in HGD, as other new approaches did before. Nonetheless, it is also obvious that NGS encompasses a multitude of “hidden costs”, which will not decline, but stay the same or even get higher with time – this should be announced clearly, as it is anyway obvious to everyone working in the field.
Thomas Liehr
Jena University Hospital, Friedrich Schiller University, Institute of Human Genetics,
Am Klinikum 1, D-07747 Jena, Germany
Publication
Human genetic diagnostics – does it always have to be NGS? (in German)
Liehr T, Weise A, Schreyer I
BIOspektrum 2017 March
Related Articles:
 | UCLA researchers develop high-sensitivity… A significant advancement for point-of-care medical diagnostics, a team of researchers from UCLA has introduced a deep learning-enhanced, paper-based vertical flow assay (VFA) capable of detecting cardiac troponin I (cTnI)… |
 | Fatigue assessment strategy using Bayesian techniques The fulfilment of the probabilistic fatigue characterization of materials based on the S-N field requires the definition of the statistical distribution of the percentile curves of the failure probability, interpreted… |
 | Rabbits with mammary carcinomas as a model for… Within a breeding colony of rabbits, the American pathologist Harry Greene (1904-1969) observed that mammary carcinomas were restricted to certain families. This is suggestive of a familiar predisposition as it… |
 | Looking inside the heart: how multiple chronic… The aim of this study was to understand how having several ongoing health problems—what we refer to as multimorbidity—impacts the heart in people with cardiovascular disease, especially those undergoing heart… |
 | Making Christmas trees under duress, or how cells… Some of the most enduring images for a molecular biologist are electron microscopy micrographs of the so-called “Christmas trees”, famously first observed by Oscar Miller from newt oocytes in 1969.… |
 | Linking copper, tyrosines and protein aggregation in… The prevalence of Alzheimer’s disease (AD), the most common cause of dementia nowadays, is expected to increase within the coming years, representing a serious threat for society and the healthcare… |







Leave a Reply
You must be logged in to post a comment.