Calmodulin is necessary for vegetative growth, ultraviolet survival, and sexual development in the model filamentous fungus Neurospora crassa
Calcium (Ca2+) is an important player of intracellular signaling system in living organisms including fungi. In the model filamentous fungus Neurospora crassa, calmodulin (CaM) is a high affinity Ca2+-binding protein involved in the Ca2+ signaling process to regulate numerous cell functions. The N. crassa CaM is highly similar to its human homologue, however, it is less acidic than its vertebrate counterpart with amino acid composition is typical to that of plant homologues. CaM is an essential gene in N. crassa, deleting the CaM gene or its direct knockout would be lethal. Therefore, in a previous study, we used CaM antagonists trifluoperazine (TFP) and chlorpromazine (CPZ) to study the functions of CaM in N. crassa.

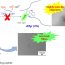
Fig. 1. Overview of CaM and Ca2+/CaMK-2 function in N. crassa. CaM is a dynamic dumbbell-shaped Ca2+-binding protein, which responds to a wide range of Ca2+ concentrations (~10-12 M -10-6 M), binding of Ca2+ (black dots) changes the CaM protein confirmation and charge. CaM bind to different target enzymes and proteins to amplify the Ca2+-signaling cascade, including a Ser/Thr kinase Ca2+/CaMK-2 (green triangle) to perform various cell functions including vegetative growth, UV survival, and sexual development.
In the Ca2+ signaling process, CaM interacts with target proteins including the conserved Ca2+/CaM-dependent Ser/Thr protein kinase 2 (Ca2+/CaMK-2). Like other Ca2+/CaMK-2 proteins, its N. crassa homologue also contains a conserved N-terminal catalytic domain and a C-terminal regulatory domain containing overlapping autoinhibitory and Ca2+/CaM binding. Binding of Ca2+/CaM displaces the autoinhibitory segment and activates the kinase, and then the kinase lock itself into activated states by autophosphorylation on a conserved threonine residue in the autoinhibitory segment that increases its affinity for CaM to about 1000-fold. We generated N. crassa strains duplicated for the CaM gene. In N. crassa, duplicated DNA sequences are the target of repeat-induced point mutation (RIP) that causes multiple G:C to A:T mutations and methylation of remaining cytosine residues within both copies of the duplicated DNA sequences after fertilization but before karyogamy during the sexual cycle. Because, N. crassa produces thousands of ascospores in the sexual cycle, it is possible to screen a large number of progeny for various RIP-induced mutations in the CaM. However, because CaM is essential, we expected to isolate only the CaMRIP strains containing viable mutations in the CaM gene. We isolated a RIP-induced CaM mutant, designated as cmdRIP(26), that vegetative phenotypes such as slow growth rate, reduced aerial hyphae development, and reduced accumulation of carotenoid that is responsible for the characteristic orange pigmentation in N. crassa, than the wild type strain. Moreover, the viability of cmdRIP(26) was severely reduced on exposures to ultraviolet (UV) irradiations. Furthermore, we found that the product of the CaM during meiosis was essential for full fertility in N. crassa. In addition, in the N. crassa Ca2+/CaMK-2 protein, we introduced S247A and T267A mutations in the candidate phosphorylation sites in the catalytic domain, and L309D mutation in the predicted CaM binding domain, and studied the phonotypes of these mutant strains. We found that the crosses homozygous for the camk-2S247A and camk-2T267A showed an intermediate phenotype (produced few hundreds of ascospores), indicating that phosphorylation of serine 247 and threonine 267 residues play a role in full fertility in N. crassa. In addition, cmd gene was upregulated in the strains lacking the Ca2+/CaMK-2, which might suggest that CaM might compensate the lack of Ca2+/CaMK-2 in N. crassa. Thus, in this study, CaM has been found necessary for normal vegetative and sexual developments, and UV survival. In addition, serine 247 and threonine 267 residues were found important for the Ca2+/CaMK-2 functions in N. crassa.
Vijya Laxmi 1,2, Ranjan Tamuli 1
1Department of Biosciences and Bioengineering, Indian Institute of Technology Guwahati, Assam, India
2Department of Agricultural Sciences, SGT University, Shree Guru Gobind Singh Tricentenary,
Gurugram, Haryana, India
Publication
The calmodulin gene in Neurospora crassa is required for normal vegetative growth, ultraviolet survival, and sexual development.
Laxmi V, Tamuli R
Arch Microbiol. 2017 May












Leave a Reply
You must be logged in to post a comment.