Mitochondrial POLG2 disease mutations impair cellular energy supply
Mitochondrial diseases are devastating disorders for which there is no cure and no proven treatment. About 1 in 2000 individuals are at risk of developing a mitochondrial disease sometime during the course of their life. Half of those affected are children who show symptoms before age 5 and approximately 80% of these will die before age 20. Mitochondrial genetic diseases are characterized by alterations in the mitochondrial genome, as point mutations, deletions, rearrangements, or depletion of the double stranded mitochondrial DNA (mtDNA). Mutations in human mtDNA cause premature aging, severe neuromuscular pathologies and maternally inherited metabolic diseases. Our mtDNA is replicated and repaired by a three subunit enzyme, DNA polymerase γ (polγ), composed of: 1) a 140-kDa catalytic subunit (p140) harboring the DNA synthesis active site, encoded by the POLG gene and 2) an ~110-kDa dimeric processivity subunit (composed of two p55 monomers) that prevents polγ from falling of the mtDNA during replication, encoded by POLG2. Other factors help polg replicate the mitochondrial genome and comprise

Fig. 1. Human mtDNA replication. A. Cartoon of the key components of the mitochondrial replisome. B. Live cell confocal fluorescence microscopy of HEK-293 cells harboring GFP-p55.
the mtDNA replication machinery or replisome (Fig. 1 A). Key components of this machinery include: 1) the topoisomerase that unwinds DNA ahead of pol g by enzymatically breaking and rejoining it allowing the DNA strands to pass through one another, 2) the Twinkle helicase that disrupts the bonds that hold the two DNA strand together thereby separating them, and 3) the mitochondrial single stranded DNA binding protein (mtSSB) that stabilizes mtDNA during replication. Nearly 300 pathogenic mutations in POLG cause a wide spectrum of disease including progressive external ophthalmoplegia (PEO), parkinsonism, premature menopause, Alpers syndrome, mitochondrial neurogastrointestinal encephalomyopathy (MNGIE) or sensory ataxic neuropathy, dysarthria, and ophthalmoparesis (SANDO). Similar to POLG, disease mutations have also been identified in the POLG2 and Twinkle genes underscoring their importance in human mtDNA replication.
Utilizing live cell confocal fluorescence microscopy we demonstrated that the p55 polγ processivity subunit can be tagged with green fluorescent protein (GFP) and retains the ability to localize to mtDNA molecules within the cell (visualized as sharp green points or dots in the cytoplasm that colocalize with DAPI-stained mtDNA blue dots, Fig. 1 B). Also, we demonstrated that the native mitochondrial targeting sequence (MTS) is required for directing p55 to the mitochondrion and if it is removed p55 remains trapped in the cytoplasm following cytoplasmic translation (Fig. 2 A, d – f). Furthermore, although P205R and L475DfsX2 p55 disease variants retain the ability to localize to mitochondria, they misfold and are unable to localize to mtDNA (Fig. 2 A).

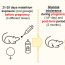
Fig. 2. Analysis of POLG2 disease mutations. A. P205R and L475DfsX2 substitutions alter p55’s mitochondrial localization. B. G451E p55 poisons polγ’s processivity via a dominant negative mechanism. C. POLG2 disease mutations impair reserve respiratory capacity, a measure of cellular energy supply
Because all currently identified POLG2 patients harbor heterozygous mutations and because monomers within the p55 dimer do not readily dissociate, we theorize that patients harbor a mixture of p55 molecules: 25% wild-type homodimers, 25% variant homodimers, and 50% heterodimers. Using a tandem affinity strategy and biochemistry to study p55 heterodimers we showed that one p55 disease variant, G451E, is dominant negative and associates with a wild-type p55 monomer in polγ to poison the enzyme’s activity (Fig. 2 B). To help explain this effect we determined that the G451E p55 heterodimer has a weakened ability to bind to p140 which prevents polγ from incorporating the normal amount of DNA building blocks (nucleotides) into newly synthesized DNA.
Lastly, cell lines harboring individual POLG2 disease mutations have reduced energy supplies (Fig. 2 C). We hypothesize that the various defects associated with p55 disease variants ultimately result in diminished cellular energy reserves and by extension mitochondrial disease. This study aids in our understanding of how these complex disorders develop and provides valuable cell line models of mitochondrial disease.
Matthew J Young and William C Copeland
Publication
POLG2 disease variants: analyses reveal a dominant negative heterodimer, altered mitochondrial localization and impaired respiratory capacity.
Young MJ, Humble MM, DeBalsi KL, Sun KY, Copeland WC
Hum Mol Genet. 2015 Sep 15
One Response to Mitochondrial POLG2 disease mutations impair cellular energy supply
Leave a Reply
You must be logged in to post a comment.












Mitochondrial diseases Seemed to be of a quite serious situation. Do hope great step can be made in the research of it. Anyway, its risk is too high.