Improved prediction of accessible surface area
Protein consists of a linear chain of amino acid residues connected by peptide bond. Protein performs an array of functions by its three dimensional (3D) shapes or structures and the shape can be determined from its one dimensional (1D) amino acid sequence. Given a 1D sequence information, to determine, protein’s 3D shape thus can be useful – however, this mapping of 1D to 3D is a challenging task. Predicted accessible surface area (ASA) of a 3D proteins can assist in determining the shape of a protein.
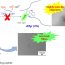
The ASA is the surface area of a biomolecule (residues) that is accessible to a spherical solvent while probing the surface of that molecule (see Figure 1). Most protein molecules have a hydrophobic core, which is not accessible to solvent and a polar surface in contact with the environment. Therefore, the ASA of amino acid residues determines the interaction pane which eventually play an important role in protein binding mechanisms, structures and functions.

Fig. 1. The dark central area, composed of atoms, can be thought of a 3D protein and the outline around that area can be thought of accessible surface area that area.
A residue can be classified as either exposed or, buried as well as can be quantified by a real value giving a more practical sense of continuously varying surface area. We attempt to predict the real valued ASA of protein residues directly from protein’s amino acid sequence.
We proposed a new sequence-based statistical predictor of ASA, namely REGAd3p. For this, we constructed a benchmark dataset, SSD1299 containing 1299 protein sequences, to train and independently test the predictor model. We generated a comprehensive set of 55 features that can reflect the intrinsic correlation between the protein sequence and the ASAs of its residues. As a part of these feature generation step, we separately build our own model for protein secondary structure (helix, beta and coil) prediction and utilized the output in the ASA prediction. We developed the ASA predictor model using regularized exact regression technique combined with genetic algorithm to optimize the weights to calculate the predicted ASA. Specifically, we supplied the features into the algorithm to learn about the appropriate characterization of ASA from the training dataset and later evaluated its performance on test dataset.
We optimized and assessed our predictor in terms of both mean absolute error (MAE) and Pearson’ correlation coefficient (PCC). We applied 10-fold cross validation as well as used independent test datasets to estimate our model’s performance reliably. The key steps in building our model are:
- We measured the performance of the predictor with and without the optimization of the weights. Optimization was done by a genetic algorithm. The predictor resulted 0.96% improved PCC and 6.14% improved MAE while optimization was included.
- We varied kernel of regularized regression algorithm from 1 to 4 to find the best parameter of the trainer. The, degree 3 polynomial function provided the best results.
- The PCC and MAE values of the final predictor of ASA, the REGAd3p, were 0.7337 and 23.9%, respectively.
- We reported case studies to depict the usefulness of the predictor.
- Our analysis showed that the predicted ASA values are quite consistent with physical properties of amino acid.
- Finally, we applied the predicted ASA on a crucial application related to the protein structure prediction. We integrated the result of ASA prediction to improve the performance of our existing energy function, namely 3DIGARS. We converted the error associated with ASA prediction into an energy component and by adding that improved our developed energy function 3DIGARS by 32.32% based on benchmark decoy test sets.
Publication
Improved prediction of accessible surface area results in efficient energy function application.
Iqbal S, Mishra A, Hoque MT
J Theor Biol. 2015 Sep 7












Leave a Reply
You must be logged in to post a comment.