Conformal invariants for landmark curve based brain morphometry analysis
Abnormal structural changes measured on cortical surfaces are important biomarkers for Alzheimer’s disease (AD). In brain imaging research, landmark curves are usually delineated on cortical surfaces to represent significant sulcus and gyrus patterns. Landmark curves are widely adopted for brain surface registration and quantitative morphometry analysis. There is increasing evidence suggesting that a landmark-based method may provide better results for cortical surface shape analysis, when compared with methods that build cortical correspondence by aligning curvature, sulcal depth or other geometric features. Furthermore, brain landmark curves are often directly analyzed to illustrate brain morphometry changes. However, most existing landmark curve based frameworks mainly focus on the landmark curves themselves. Missing at this time is a method that measures the whole brain shape changes through the analysis of landmark curves.

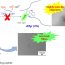
Fig. 1. A cortical surface with six landmark curves, which are automatically labeled with Caret, showing in two different views on both the original and inflated surfaces. The inflated surface is obtained with FreeSurfer.
This work presents a landmark curve based brain morphometry analysis system, based on surface conformal geometry and Teichmüller shape space theory. In practice, all oriented metric surfaces, such as the cortical surfaces, have conformal structures. Two surfaces are conformally equivalent if they can be conformally transformed to each other, i.e., there exists a smooth, revertible and angle-preserving function that converts one surface to the other. Conformal equivalent surface have the same coordinates in the Teichmüller space. The coordinates are invariant to conformal maps, so they are called conformal invariants. Conformal invariants are intrinsic features to a surface and they may serve as shape descriptors for cortical surfaces to characterize AD symptoms.
We usually model landmark curves on cortical surfaces as open boundaries. By slicing a cortical surface along the landmark curves, it becomes a multiply connected surface. Conformal invariants of multiply connected surfaces are defined as the lengths of the open boundaries in hyperbolic parameter domain and can be computed efficiently using the hyperbolic Ricci flow method. Ricci flow deforms surface metric, i.e., the unit for measuring curve lengths on a surface, with respect to surface Gaussian curvature. When the target Gaussian curvature is obtained, the resulting surface metric, which is called uniformization metric, is conformal to the original metric. When the target Gaussian curvature is negative, the uniformization metric is hyperbolic. Boundary lengths measured with the uniformization metric are the conformal invariants of the surface.
Our system consists of following steps: firstly, given an MR image, cortical surfaces are reconstructed with the Freesurfer package; secondly, six landmarks, including Central Sulcus, Anterior Half of the Superior Temporal Gyrus, Sylvian Fissure, Calcarine Sulcus, medial Wall Ventral Segment, and Medial Wall Dorsal Segment, are labeled on each cortical surface using Caret software, as shown in Figure 1; thirdly, we slice each cortical surface along its six landmarks; finally, we compute the hyperbolic uniformization metric on each multiply connected cortical surface and compute its conformal invariants.
We applied our method to study cortical shape morphometry associated with AD. We randomly selected 30 AD patients and 30 healthy people from the Alzheimer’s Disease Neuroimaging Initiative. For each cortical surface, we computed the conformal invariants as a 61 vector. Then we used the Hotelling’s T2 test to measure the difference between the two groups and permutation test was applied to correct for multiple comparisons. With conformal invariants, significant cortical shape changes due to AD were detected (p = 0.0133). We also compared with other surface features, including the Euclidean lengths of the landmark curves, cortical surface volume and surface area. None of these features revealed significant difference between the two groups, supporting the efficacy of the proposed method as a new, succinct and informative tool to study landmark curve based brain morphometry.
Jie Shi 1, Wen Zhang 1, Miao Tang 1, Richard J. Caselli 2, Yalin Wang 1
1School of Computing, Informatics, and Decision Systems Engineering,
Arizona State University, Tempe, AZ, USA
2Department of Neurology, Mayo Clinic Arizona, Scottsdale, AZ, USA
Publication
Conformal invariants for multiply connected surfaces: Application to landmark curve-based brain morphometry analysis.
Shi J, Zhang W, Tang M, Caselli RJ, Wang Y
Med Image Anal. 2017 Jan












Leave a Reply
You must be logged in to post a comment.