Human protein phosphatases: Proteomic insights
Protein phosphorylation-dephosphorylation, carried-out by kinase and phosphatases respectively, serves as major regulatory switch for controlling nearly all signaling processes. So far, the kinases remained a centerpiece in driving the functional response to signaling changes. However, more recently, the counteracting role of protein phosphatases has been much appreciated, not only in setting-up the duration and level of various signaling events but also in emergence of human diseases.

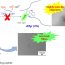
Fig. 1. Building the human protein phosphatase interaction puzzle. The figure illustrates the human protein phosphatases in the form of the most well-studied phosphatase, the protein phosphatase 2A (PP2A). PP2A is composed of the structural A and catalytic C subunits, and a regulatory B subunit. When the PP2A catalytic C subunit associates with the A and B subunits several protein isoforms, namely holoenzymes, are produced with distinct activity and substrate specificity.
About 140 protein phosphatases have been recognized in humans, dephosphorylating phospho-Serine (pS), -Threonine (pT), and -Tyrosine (pY) residues, which can be structurally assigned to three major families: Phosphoprotein phosphatase family (PPP), Protein phosphatase metal-dependent family (PPM), and Protein tyrosine family (PTP). Like other cellular enzymes and signaling proteins, phosphatases also functions as multiprotein complexes (stabilized by protein-protein interactions) and, in past few years, a significant knowledge-base has been developed to better understand their evolution, multimeric assembly, substrate recognition mechanism, and spatio-temporal regulation (Fig. 1). Despite the advancement, we are still lacking vital information on phosphatase-substrate relationships, fine-tuning phosphatase complex activity and laying foundation for phosphatase-based informational networks (phosphatase interactome).
In our study, we aimed to identify proteome level determinants of human protein phosphatase complexes (PPP, PPM, and PTP families) and decipher the activity-dependent dynamic changes of their complex composition, majorly attributed to the protein interaction remodeling. Therefore, we used affinity purification-mass spectrometry (AP-MS) based label-free systematic approach to globally search and quantify the profiles of protein phosphatase complexes. From this comprehensive characterization (54 catalytic phosphatases and 12 regulators) we found a host of novel phosphatase interactions, especially for the members of poorly represented phosphatase families (PPM and PTP), with various cellular proteins (interactors). Intriguingly, the in-depth analysis of these proteins explained the inter-relationships between various phosphatase complexes, with distinctive and non-overlapping biological functions of different phosphatase families, which certainly defied the notion of promiscuity and redundancy of phosphatase action. As exemplified by comparison, the absolute association of PTP family phosphatase interactors with extracellular region or membrane-bound vesicles in our data, clearly, emphasized their specific role in cell-surface signaling. Moreover, the display of different structural domains and motifs (HEAT, TCP1, and WD40 etc.) by these interactor proteins also expanded our view on the organizational diversity and biochemical differences of the phosphatase complexes. Another remarkable aspect of our phosphatase interactome study was the networking of phosphatases with many cancer associated proteins (CAPs). The potentially relevant links of phosphatases to particular cancer types (brain tumors and leukemias) not only signified the role of phosphatases in maintaining cellular homoeostasis, but also provided the possibly new avenues for development of phosphatase-specific therapeutics.
We further proceeded to deduce the qualitative and quantitative dynamics of the phosphatase complex assemblies by utilizing the potent phosphatase-inhibitor (PP1 and PP2A) okadaic acid (OA). Interestingly, the inhibitor-based structure-activity modulations in the interaction profiles for PPP family members, measured as the differences in the relative protein abundances (label-fee MS1 quantification), were more evident for PP1 and PP2A phosphatases, despite their highly conserved active-sites. For example, ~3-fold decrease in the abundances of several sub-constituents (striatin, integrator, and TCP1-ring proteins) of the composite PP2A complex (a catalytic and five regulatory subunits) in response to OA, indicated a noticeable reorientation of protein interactions around its enzymatic core. Further, proteomic inspection lead to identification of site-specific phosphorylation changes, which not only explained the OA-induced structural alterations but also furnished the insight on factors of phosphatase-substrate assembly or recognition and activity.
To summarize, the global analysis of human protein phosphatases enabled us to draw more complete picture of the mechanisms of phosphatase complex assembly and activity regulation, which highlighted their indispensable contribution in cell life.
Leena Yadav, Markku Varjosalo
Institute of Biotechnology, University of Helsinki, Helsinki, Finland
Publication
Systematic Analysis of Human Protein Phosphatase Interactions and Dynamics.
Yadav L, Tamene F, Göös H, van Drogen A, Katainen R, Aebersold R, Gstaiger M, Varjosalo M
Cell Syst. 2017 Apr 26












Leave a Reply
You must be logged in to post a comment.