Novel biophysical assay reveals that bacterial cell wall maintaining proteins change interaction conformation based on activity
Bacterial infections are increasingly difficult to treat with antibiotics due to antibiotic resistance. Knowledge on how of bacteria multiply needs to be profoundly expanded as a requirement for the development of new methods for the discovery of new antibiotics. Bacterial cell division may be a good antibiotic target as many of the proteins involved are unique to bacteria. The bacterium Escherichia coli is encapsulated by two lipid layers creating an inside and an outside compartment. The space between the layers is called the periplasm and contains a mesh of peptide-bridged sugars the peptidoglycan (PG) layer. The PG layer provides the bacterial cell with its shape and protects it from outside stresses. The PG layer is under constant (re)construction because it needs to facilitate the growing bacterium. The importance of the PG layer for bacterial survival and the uniqueness of the enzymes that synthesize it make it an attractive target for new antibiotics. The well-known beta-lactam antibiotics disable some of the enzymes that synthesize the PG layer. This destabilizes the PG layer causing the bacterium to eventually die. E. coli has found interesting ways to protect itself from beta-lactam antibiotics: it can inactivate the antibiotic or actively extrude it but can also build an alternative PG layer by use of enzymes that are not affected by the antibiotic. An essential step for the construction of the alternative PG layer is performed by a class of enzymes called carboxypeptidases. Previous work suggests that carboxypeptidases function by interacting as pairs called dimers and localize to sites of active PG-layer synthesis. We aimed to study their self-interaction under different activity conditions in living cells.

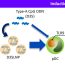
Fig. 1. Schematic overview of the Escherichia coli’s cell envelope emphasizing the periplasm. The positive and negative controls of our periplasmic FRET assay are shown as well as their rates of energy transfer. The assayed carboxypeptidase self-interactions (boxed) show the increase in energy transfer rates upon inactivating by substrate availability or mutation. The inhibited interactions show more energy transfer regardless of whether the carboxypeptidases are inactive because no substrate is available or they are made inactive by a mutation.
The preferred method to measure protein-interactions in living cells is Förster Resonance Energy Transfer (FRET). FRET requires two compatible fluorescent proteins (FP)s to be in such close proximity that energy of one (donor) is transferred to the other (acceptor). FPs can be conveniently used to tag any protein of interest, which then becomes detectable by fluorescence. If FRET is detected for two tagged proteins of interest, they are very close and are likely interacting.
FRET assays were not available for the periplasm since most FPs do not work in the periplasm. Because this is where the carboxypeptidases reside, we had to develop an assay ourselves. We found that the expression of FPs in the periplasm causes toxic effects for the cells at high concentrations. However, a red FP that was known to fluoresce in the periplasm, mCherry (mCh), could be expressed with limited toxicity to the cells. The green FP mNeonGreen (mNG) structurally resembles mCherry and appeared to fluorescence in the periplasm at reduced toxicity. The biophysical properties of mNG and mCh are identical when expressed either in the periplasm or in the interior of the cell. mNG and mCh were used together in a FRET assay in living cells showing activity-based differences in self-interaction of the main carboxypeptidases of E. coli. This was interesting as these enzymes are structurally very similar and were thought to have a redundant functionality. Inactivating the carboxypeptidases by mutation showed an increase in FRET, suggesting that they have a different conformation because likely the distance between the FPs had shortened. Forcing inactivity of wild-type carboxypeptidases by taking away their building-block substrate (by inhibition of upstream enzymes) confirmed this hypothesis. Additionally, we observed that expressing inactive carboxypeptidases results in misshapen cells whereas cells looked normal if the active variants were just deleted. With that we showed that inactivity of carboxypeptidases is not the same as their absence suggesting more variation in functionality. With our work we have provided the scientific community with the first periplasmic FRET assay and applied it to demonstrate activity-related conformational changes of carboxypeptidases, providing new insights in how bacteria multiply.
Nils Y. Meiresonne, Tanneke den Blaauwen
Bacterial Cell Biology & Physiology, Swammerdam Institute for Life Sciences,
University of Amsterdam, Science park 904, 1098 XH Amsterdam, The Netherlands
Publication
Activity-Related Conformational Changes in d,d-Carboxypeptidases Revealed by In Vivo Periplasmic Förster Resonance Energy Transfer Assay in Escherichia coli.
Meiresonne NY, van der Ploeg R, Hink MA, den Blaauwen T
MBio. 2017 Sep 12












Leave a Reply
You must be logged in to post a comment.