Visualizing transcription factor dynamics, one molecule at a time
In a multicellular organism, each cell type expresses only a subset of genes encoded in the genome. Moreover, hundreds of genes can be regulated by external stimuli. This complex regulation is mediated by proteins called transcription factors. This special type of proteins can bind to specific sequences in the genome called response elements, within regulatory regions known as enhancers. Upon binding, transcription factors serve as a “scaffold” to recruit several other proteins such as cofactors and chromatin remodelers that would ultimately promote the up- (or down) regulation of genes.

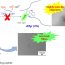
Fig. 1. Single molecules of the glucocorticoid receptor in living cells. Each green dot represents a single molecule. The nucleus (in blue) is delimited by the white-dashed line. Scale bar 6 µm.
For decades, it was thought that transcription factors stay bound to chromatin for long periods of time, allowing the sequential and well-ordered recruitment of the entire transcriptional machine. However, with the advent of live-cell fluorescence microscopy imaging, it has been demonstrated that most of transcription factors interact very transiently with DNA, in the order of just a few seconds. This radical change in paradigm raised several questions about the dynamic nature of gene expression regulation, which is still largely unresolved.
New technological advances in organic fluorophore development, microscopy techniques, and camera sensitivity have allowed researchers to visualize individual protein molecules in live cells with an unprecedent spatial and temporal resolution. This allows us to directly quantify the time each individual transcription factor remain bound to chromatin. In our study, we describe the methodology used to visualize and track the dynamics of transcription factors at the single-molecule level (Single-molecule Tracking, SMT), and further discuss the current challenges this exciting new field faces.
To be able to visualize and follow individual fluorescent proteins, it is imperative to work with the brightest, more stable fluorophore available. Unfortunately, genetically encoded fluorescent proteins such as GFP are too dim, and bleach to rapidly, to be of use. A better alternative employs organic fluorophore dyes, including reagents developed by the Lavis Lab at Janelia Farms. These dyes can be irreversibly bound to small proteins with high affinity binding pockets (HaloTag, SNAP-tag and CLIP-tag). By making fusion chimeras between the tags and the protein of interest, it is possible to visualize a specific transcription factor with a bright and relatively stable fluorophore. Interestingly, we found that the same dye can have different photostability behavior depending on which tag protein they are attached to. Unfortunately, this raises some concerns regarding the possibility of combining multiple tags for multi-color applications.
From the optics perspective, to capture the light emanating from a single-molecule, and discriminate it from background, requires the reduction of out-of-focus illumination. This can be achieved using “highly inclined and laminated optical sheet illumination” (HiLO), developed by the Sakata-Sogawa lab. In this special imaging mode, a laser beam hits the sample from the side instead as from below, the most common configuration. This generates a thin, oblique layer of illumination across the cell nucleus. The caveat of this type of illumination is that photobleaching is not homogenous within cells, generating possible artifacts during the data analysis. The best way to analyze SMT data is a matter of continuous debate on this relatively new field. Nevertheless, all evidence derived from SMT thus far confirms the general idea that transcription factors are highly dynamic, and only a small fraction are specifically bound to response elements at any given time. Which of these molecules are functionally active, i.e. able to recruit RNA polymerase II, is still an open question that occupies many investigators currently working in this field.
Diego M. Presman, Gordon L. Hager
Laboratory of Receptor Biology and Gene Expression, National Cancer Institute,
National Institutes of Health, Bethesda, Maryland, USA
Publication
Quantifying transcription factor binding dynamics at the single-molecule level in live cells.
Presman DM, Ball DA, Paakinaho V, Grimm JB, Lavis LD, Karpova TS, Hager GL
Methods. 2017 Jul 1












Leave a Reply
You must be logged in to post a comment.