The molecular analysis of tumor cells in pancreatic tissue – searching for a needle in a haystack
In cancer research usually the tumor cell specific abnormalities are of interest: gene mutations, production of abnormal proteins, production of abnormal quantity of proteins, tumor cells treatment response. But the molecular analysis of pancreatic tumor samples poses certain problems, which can lead to a dramatic misinterpretation of scientific results.
One problem is due to the complex composition of the pancreatic tumor sample itself. The tumor sample does not consist for the most part of tumor cells. The tumor growth is accompanied with the dramatic overgrowth of connective tissue and an immense infiltration of immune cells. As well, the tumor sample can comprise remained healthy pancreas part. This includes acinar cells, which produce the pancreas juice playing a crucial role in digesting and absorbing food and drinks and islet cells, which secrete hormones into the bloodstream. Furthermore, adipocytes (fat cells) can be incorporated in the tumor sample.
The risk of data misinterpretation is high. For this known reason, standardly a histological observation of the tissue sample is performed by a pathologist to determine the tumor cell content in the sample. But in various cases a histological observation is not feasible due the following reasons: sample is too small, sample was already homogenated, RNA or proteins were already extracted from the tissue, or, the analysis bases on expression data of online databases, where the information of the sample composition is not available.
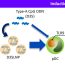
A standard method to determine tissue composition on molecular level and to stratify samples does not yet exist. Therefore, we established a workflow that we called RNACellStrat. The procedure bases on the definition of the activity of 5 genes (RNA expression analysis). We could show that these genes are specifically expressed only in one pancreatic compartment and that they can be used to estimate the composition of the sample piece (see Figure).

The RNACellStrat procedure. By analyzing the activity of 5 marker genes, we can draw conclusion to the initial composition of the pancreatic sample piece with the aim to exclude conspicuous tissue samples from further analysis.
To demonstrate the importance of the established procedure, we performed an exemplary patient survival analysis. With this statistic test we compared the survival times of patients after pancreatic cancer resection. The patients were divided into the two groups according the activity of the cancer-specific gene named S100A2. From literature, it is known that patients with S100A2-negative tumors survive longer. As starting material we randomly choose pancreatic “tumor samples” of patients with the diagnosis of PDAC (pancreatic ductal adenocarcinoma) from a large biobank. In a first step, we decided not to perform the RNAStratCell procedure before analysis. The result was that none of the two groups (S100A2-low tumors vs. S100A2-high tumors) had a better survival time. In the second step, we performed RNACellStrat procedure and excluded about one quarter of the tissue samples prior survival analysis due to their conspicuous gene activities of the 5 marker genes. The result showed a significantly longer survival (about one year) of the patients with S100A2-low tumors.
This clearly shows that an insufficient quality control of pancreatic cancer tissue samples prior to prognostic analysis can lead to wrong conclusion. Under- as well as overinterpretations may occur.
This strategy for cellular annotation is specially adapted to pancreatic tumor tissue samples, but may be transferred to other diseases and tissue types.
Dr. Anette Heller
Department of Surgery, University Hospital Heidelberg, Heidelberg, Germany
Publication
Stratification of pancreatic tissue samples for molecular studies: RNA-based cellular annotation procedure.
Heller A, Gaida MM, Männle D, Giese T, Scarpa A, Neoptolemos JP, Hackert T, Strobel O, Hoheisel JD, Giese NA, Bauer AS
Pancreatology. 2015 Jul-Aug












Leave a Reply
You must be logged in to post a comment.