New morphology of amyloid fibrils
It has been shown earlier that one of the proteins from the Saccharomyces cerevisiae cell wall (glucantransferase Bgl2) forms amyloid structures after isolation from cell wall under acid pH conditions. It is known that the acid treatment of yeast cells leads to the occurrence of an apoptotic phenotype and induction of general stress response pathways that may include the mechanisms of cell wall integrity control. The capacity of potentially amyloidogenic regions in the Bgl2p sequence to generate amyloid formation was examined using the bioinformatics analysis.

Fig. 1. EM images of fragments of fields of the AspNB preparation. Sample (0.4 mg/ml) in 5% acetic acid (5% DMSO) and incubation at 37 °C during 30 h: (A) lateral association of fibrils in the form of a wide ribbon; (B) two fibrils associated by their side surfaces; (C) two fibrils contact the Formvar support by their side surfaces; (D) fibrils in the form of bundles; (E, F) fragments of fibrils in the form of bundles. Ring oligomers associate randomly (E, arrow).
The analysis of potentially amyloidogenic regions capacity to form amyloids and understanding of the dependence of amyloid properties on different conditions (synthesis, pH, temperature, ionic strength, etc.) are important for the understanding of general mechanisms of amyloidogenesis in proteins. The process of amyloid formation was investigated for amyloidogenic fragment AspNB predicted in Bgl2 (residues 166–175, VDSWNVLVAG with unblock termini) and homologous peptide with a substitution of aspartate for glutamate in position 2 GluNB (VESWNVLVAG) by different methods. The comparison of the data, obtained for the two peptides, allowed us to conclude that substitutions of D for E affected the process of amyloid formation: AspNB forms amyloid fibrils faster than GluNB. However, such mutation has no effect on the morphology of mature amyloid fibrils.
Detailed analysis of the morphology of fibrils of the both preparations revealed strong polymorphism of fibrils (Fig. 1). Peptides form ribbons of different widths, twisted ribbons of different periods, bundles of fibrils with different diameters. Moreover, we found new morphology fibrils not mentioned in the literature (Fig. 2, С4). This morphology resembles snakes lying side by side in the form of a wave without intertwining.
The most essential result from the EM studies is that in spite of strong polymorphism of fibrils, in all cases the main building element of a fibril of any morphology is a ring oligomer with the external diameter of about 6–8 nm, the internal diameter (of the hole) of about 2–3 nm and the height of about 3–4 nm (estimated by the bending sites of a single fibril (Fig. 1)). Such oligomers associate with each other either ring to ring or slightly overlaying each other.
The data from X-ray analysis show that fibrils formed from ring oligomers have a cross-β structure. This is consistent with our proposed model of oligomer structure and the suggested mechanism of packing of oligomers in fibrils. According to the proposed model, the ring-like oligomer includes β-strands as well as 12 β-sheets. This quantity is sufficient to produce reflections characteristic of cross-β structure.
According to a simplified scheme, the formation of fibrils proceeds as follows: a monomer → a destabilized monomer → an oligomer → a fibril. The weakest point in this scheme is the oligomer. For many amyloidogeneic proteins or peptides, oligomers can be observed at the initial stage of fibril formation; oligomers (Aβ peptides, insulin) frequently have ring morphology. It is unclear what occurs at the stage ‘an oligomer → a fibril’. In what way are fibrils formed from ring-like oligomers? We believe the oligomer is quite probable to be the basic unit of a fibril. Ring-like oligomers can form any morphology of a fibril, i.e. under certain conditions polymorphism of fibrils of the same preparation can be explained by different ways of interaction of oligomers (Fig. 2). It is simpler to explain the changes in morphology of fibrils even upon changes in different conditions of fibril formation (pH, ionic strength, temperature etc.).

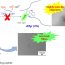
Fig. 2. Schematic representation of the fibril polymorphism. (A) Monomers; (B) oligomers; (C) single fibril. The formation from oligomers: 1) ribbons; 2) thin bundles; 3) large diameter bundles; 4) new morphology of amyloid fibrils. (D) Molecular model of stacking of ring oligomers in fibrils (top and side views); (E) the corresponding EM images.
It can be assumed that Bgl2p is also conformationally labile. This assumption is confirmed by the disclosed ability of Bgl2p to form fibrils of various morphologies (nets or bundles) under different conditions (in cultural media or in water after isolation from the cell wall).
Olga M. Selivanova 1, Alexey K. Surin 1, Tatyana S. Kalebina 2, Oxana V. Galzitskaya 1
1Institute of Protein Research, Russian Academy of sciences
2Department of Molecular Biology, Faculty of Biology, Moscow State University
Publication
Structural model of amyloid fibrils for amyloidogenic peptide from Bgl2p-glucantransferase of S. cerevisiae cell wall and its modifying analog. New morphology of amyloid fibrils.
Selivanova OM, Glyakina AV, Gorbunova EY, Mustaeva LG, Suvorina MY, Grigorashvili EI, Nikulin AD, Dovidchenko NV, Rekstina VV, Kalebina TS, Surin AK, Galzitskaya OV
Biochim Biophys Acta. 2016 Nov












Leave a Reply
You must be logged in to post a comment.