Computational refinement and validation protocol for proteins
HIV protects itself using a membrane layer of lipids which covers all proteins which could potentially be targeted by drugs. The only exposed proteins, gp120 and gp41 form spikes that protrude out of the lipid membrane. These proteins play a major role in HIV gaining entry into a cell.

Fig. 1. Structures of variable regions V1-V2, V3 are could not be determined by existing imaging protocols. However, they are very close to the binding locations of CD4 and 17b and is expected to influence the binding.
First, gp120 binds (attaches itself) to CD4, a protein which is found on the cellular membrane of human T-cells. Then, gp120 undergoes a change of structure or configuration, which allows it to bind with another membrane protein, CCR5. Finally, these interactions allow gp41 to change its configuration which pulls the membrane off HIV and allows it to fuse with the T-cell lipid membrane. To understand and disrupt this entry mechanism of HIV into cells, we need to know atomic level details of gp120 and its functional interaction with CD4 and CCR5.
However, the highly variable and flexible nature of gp120 has prevented current imaging methods from being able to completely resolve the atomic structure gp120. gp120 have large variable regions (segments of the protein that are highly flexible), which are suspected to be important parts of the binding interaction. See Figure 1.

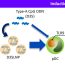
Fig. 2. [Left top] Previous known model of the interaction of gp120 with CD4 and 17b. Note that each of them appears in three copies, since the HIV spikes are trimers of gp120 and gp41. [Bottom left] The new model which includes the variable regions of gp120 as well as the configuration of partial fragments of gp41 in bound configuration. [Right] A close-up showing 17b, CD4 and gp41 as cartoon ribbons, and highlighting the variable regions of gp120 using different colors. The interaction of the V1-V2 region with CD4, and the V3 region with 17b is evident.
The model showed (Fig. 2), in particular, how the V1-V2 variable regions are expected to move, from its unbound state to allow binding of CD4. We also found that the V3 region, and to a small extent the V1-V2 regions also move to allow binding of 17b (a mimic of CCR5), thus gaining new insights into details of the HIV – T4 cell entry mechanism.
Publication
Computational Refinement and Validation Protocol for Proteins with Large Variable Regions Applied to Model HIV Env Spike in CD4 and 17b Bound State.
Rasheed M, Bettadapura R, Bajaj C.
Structure. 2015 Jun 2












Leave a Reply
You must be logged in to post a comment.